Note
Click here to download the full example code
EEG forward operator with a template MRI¶
This tutorial explains how to compute the forward operator from EEG data
using the standard template MRI subject fsaverage.
Caution
Source reconstruction without an individual T1 MRI from the subject will be less accurate. Do not over interpret activity locations which can be off by multiple centimeters.
This tutorial covers:
# Authors: Alexandre Gramfort <alexandre.gramfort@inria.fr>
# Joan Massich <mailsik@gmail.com>
#
# License: BSD Style.
import os.path as op
import mne
from mne.datasets import eegbci
from mne.datasets import fetch_fsaverage
# Download fsaverage files
fs_dir = fetch_fsaverage(verbose=True)
subjects_dir = op.dirname(fs_dir)
# The files live in:
subject = 'fsaverage'
trans = 'fsaverage' # MNE has a built-in fsaverage transformation
src = op.join(fs_dir, 'bem', 'fsaverage-ico-5-src.fif')
bem = op.join(fs_dir, 'bem', 'fsaverage-5120-5120-5120-bem-sol.fif')
Out:
0 files missing from /home/circleci/project/mne/datasets/_fsaverage/root.txt in /home/circleci/mne_data/MNE-fsaverage-data
0 files missing from /home/circleci/project/mne/datasets/_fsaverage/bem.txt in /home/circleci/mne_data/MNE-fsaverage-data/fsaverage
Load the data¶
We use here EEG data from the BCI dataset.
Note
See Plotting sensor layouts of EEG systems to view all the standard EEG montages available in MNE-Python.
raw_fname, = eegbci.load_data(subject=1, runs=[6])
raw = mne.io.read_raw_edf(raw_fname, preload=True)
# Clean channel names to be able to use a standard 1005 montage
new_names = dict(
(ch_name,
ch_name.rstrip('.').upper().replace('Z', 'z').replace('FP', 'Fp'))
for ch_name in raw.ch_names)
raw.rename_channels(new_names)
# Read and set the EEG electrode locations
montage = mne.channels.make_standard_montage('standard_1005')
raw.set_montage(montage)
raw.set_eeg_reference(projection=True) # needed for inverse modeling
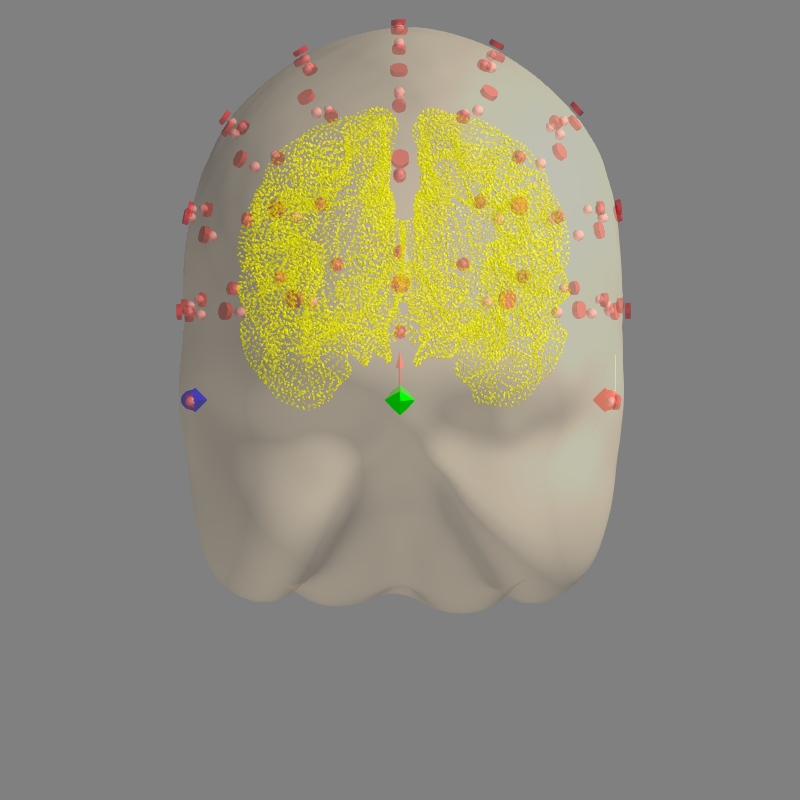
# Check that the locations of EEG electrodes is correct with respect to MRI
mne.viz.plot_alignment(
raw.info, src=src, eeg=['original', 'projected'], trans=trans,
show_axes=True, mri_fiducials=True, dig='fiducials')
Out:
Extracting EDF parameters from /home/circleci/mne_data/MNE-eegbci-data/files/eegmmidb/1.0.0/S001/S001R06.edf...
EDF file detected
Setting channel info structure...
Creating raw.info structure...
Reading 0 ... 19999 = 0.000 ... 124.994 secs...
Adding average EEG reference projection.
1 projection items deactivated
Average reference projection was added, but has not been applied yet. Use the apply_proj method to apply it.
Reading /home/circleci/mne_data/MNE-fsaverage-data/fsaverage/bem/fsaverage-ico-5-src.fif...
Using outer_skin.surf for head surface.
Setup source space and compute forward¶
fwd = mne.make_forward_solution(raw.info, trans=trans, src=src,
bem=bem, eeg=True, mindist=5.0, n_jobs=1)
print(fwd)
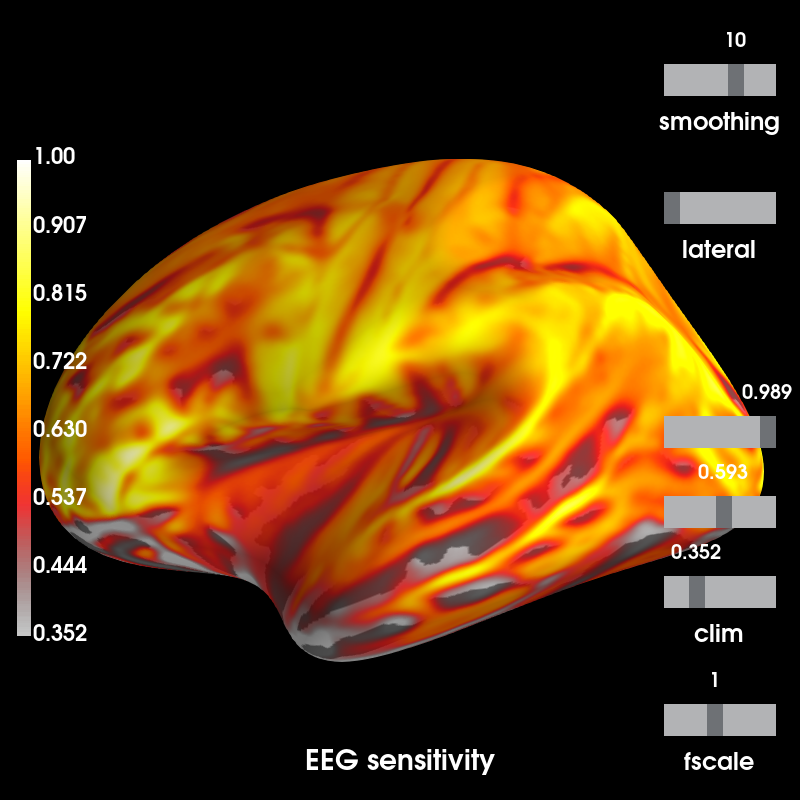
# Use fwd to compute the sensitivity map for illustration purposes
eeg_map = mne.sensitivity_map(fwd, ch_type='eeg', mode='fixed')
brain = eeg_map.plot(time_label='EEG sensitivity', subjects_dir=subjects_dir,
clim=dict(lims=[5, 50, 100]))
Out:
Source space : /home/circleci/mne_data/MNE-fsaverage-data/fsaverage/bem/fsaverage-ico-5-src.fif
MRI -> head transform : /home/circleci/project/mne/data/fsaverage/fsaverage-trans.fif
Measurement data : instance of Info
Conductor model : /home/circleci/mne_data/MNE-fsaverage-data/fsaverage/bem/fsaverage-5120-5120-5120-bem-sol.fif
Accurate field computations
Do computations in head coordinates
Free source orientations
Reading /home/circleci/mne_data/MNE-fsaverage-data/fsaverage/bem/fsaverage-ico-5-src.fif...
Read 2 source spaces a total of 20484 active source locations
Coordinate transformation: MRI (surface RAS) -> head
0.999994 0.003552 0.000202 -1.76 mm
-0.003558 0.998389 0.056626 31.09 mm
-0.000001 -0.056626 0.998395 39.60 mm
0.000000 0.000000 0.000000 1.00
Read 64 EEG channels from info
Head coordinate coil definitions created.
Source spaces are now in head coordinates.
Setting up the BEM model using /home/circleci/mne_data/MNE-fsaverage-data/fsaverage/bem/fsaverage-5120-5120-5120-bem-sol.fif...
Loading surfaces...
Loading the solution matrix...
Three-layer model surfaces loaded.
Loaded linear_collocation BEM solution from /home/circleci/mne_data/MNE-fsaverage-data/fsaverage/bem/fsaverage-5120-5120-5120-bem-sol.fif
Employing the head->MRI coordinate transform with the BEM model.
BEM model fsaverage-5120-5120-5120-bem-sol.fif is now set up
Source spaces are in head coordinates.
Checking that the sources are inside the surface and at least 5.0 mm away (will take a few...)
Skipping interior check for 2433 sources that fit inside a sphere of radius 47.7 mm
Skipping solid angle check for 0 points using Qhull
Skipping interior check for 2241 sources that fit inside a sphere of radius 47.7 mm
Skipping solid angle check for 0 points using Qhull
Setting up for EEG...
Computing EEG at 20484 source locations (free orientations)...
Finished.
<Forward | MEG channels: 0 | EEG channels: 64 | Source space: Surface with 20484 vertices | Source orientation: Free>
64 out of 64 channels remain after picking
Adding average EEG reference projection.
Using control points [0.35186294 0.5925817 1. ]
Total running time of the script: ( 0 minutes 32.460 seconds)
Estimated memory usage: 331 MB