Note
Click here to download the full example code
Generate simulated evoked data¶
Use simulate_sparse_stc() to simulate evoked data.
# Author: Daniel Strohmeier <daniel.strohmeier@tu-ilmenau.de>
# Alexandre Gramfort <alexandre.gramfort@inria.fr>
#
# License: BSD (3-clause)
import numpy as np
import matplotlib.pyplot as plt
import mne
from mne.datasets import sample
from mne.time_frequency import fit_iir_model_raw
from mne.viz import plot_sparse_source_estimates
from mne.simulation import simulate_sparse_stc, simulate_evoked
print(__doc__)
Load real data as templates
data_path = sample.data_path()
raw = mne.io.read_raw_fif(data_path + '/MEG/sample/sample_audvis_raw.fif')
proj = mne.read_proj(data_path + '/MEG/sample/sample_audvis_ecg-proj.fif')
raw.info['projs'] += proj
raw.info['bads'] = ['MEG 2443', 'EEG 053'] # mark bad channels
fwd_fname = data_path + '/MEG/sample/sample_audvis-meg-eeg-oct-6-fwd.fif'
ave_fname = data_path + '/MEG/sample/sample_audvis-no-filter-ave.fif'
cov_fname = data_path + '/MEG/sample/sample_audvis-cov.fif'
fwd = mne.read_forward_solution(fwd_fname)
fwd = mne.pick_types_forward(fwd, meg=True, eeg=True, exclude=raw.info['bads'])
cov = mne.read_cov(cov_fname)
info = mne.io.read_info(ave_fname)
label_names = ['Aud-lh', 'Aud-rh']
labels = [mne.read_label(data_path + '/MEG/sample/labels/%s.label' % ln)
for ln in label_names]
Out:
Opening raw data file /home/circleci/mne_data/MNE-sample-data/MEG/sample/sample_audvis_raw.fif...
Read a total of 3 projection items:
PCA-v1 (1 x 102) idle
PCA-v2 (1 x 102) idle
PCA-v3 (1 x 102) idle
Range : 25800 ... 192599 = 42.956 ... 320.670 secs
Ready.
Read a total of 6 projection items:
ECG-planar-999--0.200-0.400-PCA-01 (1 x 203) idle
ECG-planar-999--0.200-0.400-PCA-02 (1 x 203) idle
ECG-axial-999--0.200-0.400-PCA-01 (1 x 102) idle
ECG-axial-999--0.200-0.400-PCA-02 (1 x 102) idle
ECG-eeg-999--0.200-0.400-PCA-01 (1 x 59) idle
ECG-eeg-999--0.200-0.400-PCA-02 (1 x 59) idle
Reading forward solution from /home/circleci/mne_data/MNE-sample-data/MEG/sample/sample_audvis-meg-eeg-oct-6-fwd.fif...
Reading a source space...
Computing patch statistics...
Patch information added...
Distance information added...
[done]
Reading a source space...
Computing patch statistics...
Patch information added...
Distance information added...
[done]
2 source spaces read
Desired named matrix (kind = 3523) not available
Read MEG forward solution (7498 sources, 306 channels, free orientations)
Desired named matrix (kind = 3523) not available
Read EEG forward solution (7498 sources, 60 channels, free orientations)
MEG and EEG forward solutions combined
Source spaces transformed to the forward solution coordinate frame
364 out of 366 channels remain after picking
366 x 366 full covariance (kind = 1) found.
Read a total of 4 projection items:
PCA-v1 (1 x 102) active
PCA-v2 (1 x 102) active
PCA-v3 (1 x 102) active
Average EEG reference (1 x 60) active
Read a total of 4 projection items:
PCA-v1 (1 x 102) active
PCA-v2 (1 x 102) active
PCA-v3 (1 x 102) active
Average EEG reference (1 x 60) active
Generate source time courses from 2 dipoles and the correspond evoked data
times = np.arange(300, dtype=np.float64) / raw.info['sfreq'] - 0.1
rng = np.random.RandomState(42)
def data_fun(times):
"""Function to generate random source time courses"""
return (50e-9 * np.sin(30. * times) *
np.exp(- (times - 0.15 + 0.05 * rng.randn(1)) ** 2 / 0.01))
stc = simulate_sparse_stc(fwd['src'], n_dipoles=2, times=times,
random_state=42, labels=labels, data_fun=data_fun)
Generate noisy evoked data
picks = mne.pick_types(raw.info, meg=True, exclude='bads')
iir_filter = fit_iir_model_raw(raw, order=5, picks=picks, tmin=60, tmax=180)[1]
nave = 100 # simulate average of 100 epochs
evoked = simulate_evoked(fwd, stc, info, cov, nave=nave, use_cps=True,
iir_filter=iir_filter)
Out:
Average patch normals will be employed in the rotation to the local surface coordinates....
Converting to surface-based source orientations...
[done]
Projecting source estimate to sensor space...
[done]
4 projection items deactivated
Created an SSP operator (subspace dimension = 4)
4 projection items activated
SSP projectors applied...
Plot
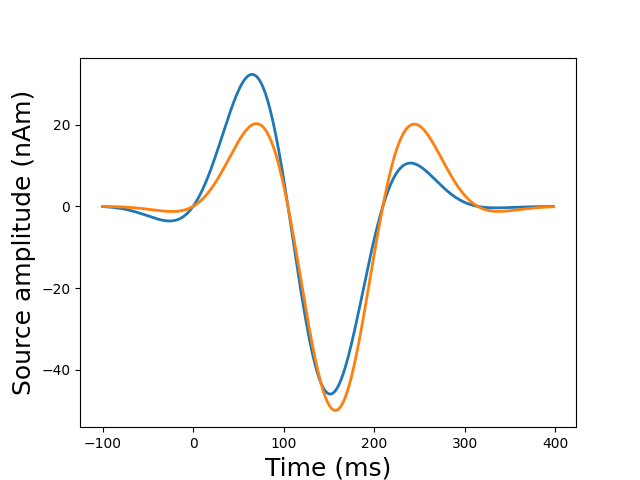
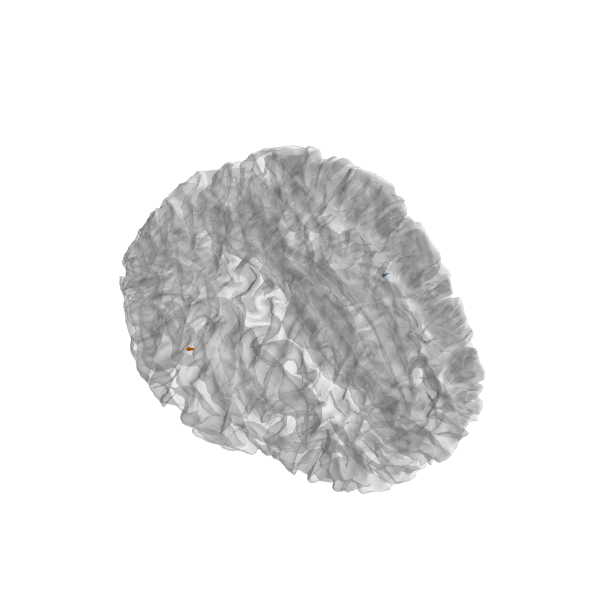
plot_sparse_source_estimates(fwd['src'], stc, bgcolor=(1, 1, 1),
opacity=0.5, high_resolution=True)
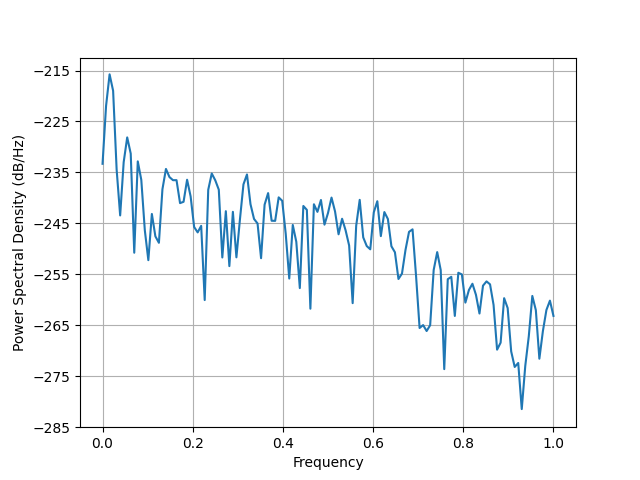
plt.figure()
plt.psd(evoked.data[0])
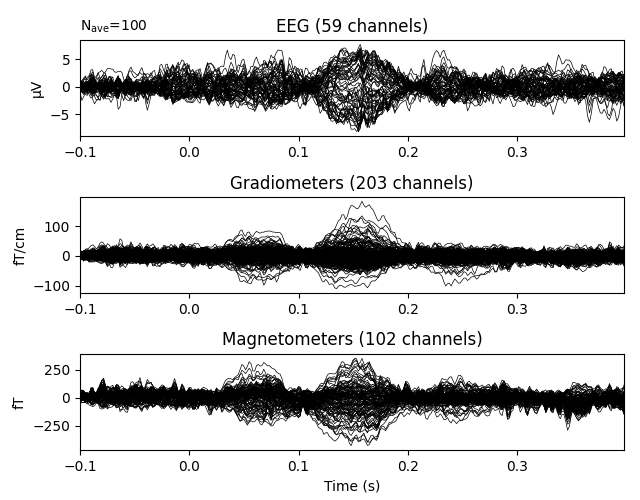
evoked.plot(time_unit='s')

Out:
Total number of active sources: 2
Total running time of the script: ( 0 minutes 7.294 seconds)
Estimated memory usage: 327 MB