Note
Click here to download the full example code or to run this example in your browser via Binder
Save and load T1-weighted MRI scan along with anatomical landmarks in BIDS¶
When working with MEEG data in the domain of source localization, we usually have to deal with aligning several coordinate systems, such as the coordinate systems of …
the head of a study participant
the recording device (in the case of MEG)
the anatomical MRI scan of a study participant
The process of aligning these frames is also called coregistration, and is
performed with the help of a transformation matrix, called trans in MNE.
In this tutorial, we show how MNE-BIDS can be used to save a T1 weighted
MRI scan in BIDS format, and to encode all information of the trans object
in a BIDS compatible way.
Finally, we will automatically reproduce our trans object from a BIDS
directory.
See the documentation pages in the MNE docs for more information on source alignment and coordinate frames
Note
For this example you will need to install matplotlib and
nilearn on top of your usual mne-bids installation.
# Authors: Stefan Appelhoff <stefan.appelhoff@mailbox.org>
# License: BSD (3-clause)
We are importing everything we need for this example:
import os.path as op
import shutil as sh
import numpy as np
import matplotlib.pyplot as plt
from nilearn.plotting import plot_anat
import mne
from mne.datasets import sample
from mne.source_space import head_to_mri
from mne_bids import (write_raw_bids, make_bids_basename, write_anat,
get_head_mri_trans)
from mne_bids.utils import print_dir_tree
We will be using the MNE sample data and write a basic BIDS dataset. For more information, you can checkout the respective example.
data_path = sample.data_path()
event_id = {'Auditory/Left': 1, 'Auditory/Right': 2, 'Visual/Left': 3,
'Visual/Right': 4, 'Smiley': 5, 'Button': 32}
raw_fname = op.join(data_path, 'MEG', 'sample', 'sample_audvis_raw.fif')
events_data = op.join(data_path, 'MEG', 'sample', 'sample_audvis_raw-eve.fif')
output_path = op.abspath(op.join(data_path, '..', 'MNE-sample-data-bids'))
if op.exists(output_path):
sh.rmtree(output_path)
raw = mne.io.read_raw_fif(raw_fname)
sub = '01'
ses = '01'
task = 'audiovisual'
run = '01'
bids_basename = make_bids_basename(subject=sub, session=ses, task=task,
run=run)
write_raw_bids(raw, bids_basename, output_path, events_data=events_data,
event_id=event_id, overwrite=True)
# Print the directory tree
print_dir_tree(output_path)
Out:
Opening raw data file /home/stefanappelhoff/mne_data/MNE-sample-data/MEG/sample/sample_audvis_raw.fif...
Read a total of 3 projection items:
PCA-v1 (1 x 102) idle
PCA-v2 (1 x 102) idle
PCA-v3 (1 x 102) idle
Range : 25800 ... 192599 = 42.956 ... 320.670 secs
Ready.
Current compensation grade : 0
Opening raw data file /home/stefanappelhoff/mne_data/MNE-sample-data/MEG/sample/sample_audvis_raw.fif...
Read a total of 3 projection items:
PCA-v1 (1 x 102) idle
PCA-v2 (1 x 102) idle
PCA-v3 (1 x 102) idle
Range : 25800 ... 192599 = 42.956 ... 320.670 secs
Ready.
Current compensation grade : 0
Creating folder: /home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg
Writing '/home/stefanappelhoff/mne_data/MNE-sample-data-bids/participants.tsv'...
participant_id age sex
sub-01 n/a n/a
Writing '/home/stefanappelhoff/mne_data/MNE-sample-data-bids/participants.json'...
{
"participant_id": {
"Description": "Unique participant identifier"
},
"age": {
"Description": "Age of the participant at time of testing",
"Units": "years"
},
"sex": {
"Description": "Biological sex of the participant",
"Levels": {
"F": "female",
"M": "male"
}
}
}
Writing '/home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/sub-01_ses-01_scans.tsv'...
filename acq_time
meg/sub-01_ses-01_task-audiovisual_run-01_meg.fif 2002-12-03T20:01:10
Writing '/home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg/sub-01_ses-01_coordsystem.json'...
{
"MEGCoordinateSystem": "Elekta",
"MEGCoordinateUnits": "m",
"HeadCoilCoordinates": {
"NAS": [
3.725290298461914e-09,
0.10260561108589172,
4.190951585769653e-09
],
"LPA": [
-0.07137660682201385,
0.0,
5.122274160385132e-09
],
"RPA": [
0.07526767998933792,
0.0,
5.587935447692871e-09
],
"coil1": [
0.032922741025686264,
0.09897983074188232,
0.07984329760074615
],
"coil2": [
-0.06998106092214584,
0.06771647930145264,
0.06888450682163239
],
"coil3": [
-0.07260829955339432,
-0.02086828649044037,
0.0971473976969719
],
"coil4": [
0.04996863007545471,
-0.007233052980154753,
0.1228904277086258
]
},
"HeadCoilCoordinateSystem": "RAS",
"HeadCoilCoordinateUnits": "m"
}
Writing '/home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg/sub-01_ses-01_task-audiovisual_run-01_events.tsv'...
onset duration trial_type value sample
3.6246181587150867 0.0 Auditory/Right 2 2177
4.237323479067476 0.0 Visual/Left 3 2545
4.946596485779753 0.0 Auditory/Left 1 2971
5.692498614904401 0.0 Visual/Right 4 3419
6.41342634238425 0.0 Auditory/Right 2 3852
Writing '/home/stefanappelhoff/mne_data/MNE-sample-data-bids/dataset_description.json'...
{
"Name": " ",
"BIDSVersion": "1.2.0"
}
/home/stefanappelhoff/Desktop/bids/mne-bids/mne_bids/write.py:407: UserWarning: No line frequency found, defaulting to 50 Hz
warn('No line frequency found, defaulting to 50 Hz')
Reading 0 ... 166799 = 0.000 ... 277.714 secs...
/home/stefanappelhoff/miniconda3/envs/mne_bids3/lib/python3.6/site-packages/mne/utils/docs.py:830: DeprecationWarning: Function read_montage is deprecated; ``read_montage`` is deprecated and will be removed in v0.20. Please use ``read_dig_fif``, ``read_dig_egi``, ``read_custom_montage``, or ``read_dig_captrack`` to read a digitization based on your needs instead; or ``make_standard_montage`` to create ``DigMontage`` based on template; or ``make_dig_montage`` to create a ``DigMontage`` out of np.arrays
warnings.warn(msg, category=DeprecationWarning)
/home/stefanappelhoff/miniconda3/envs/mne_bids3/lib/python3.6/site-packages/mne/utils/docs.py:813: DeprecationWarning: Class Montage is deprecated; Montage class is deprecated and will be removed in v0.20. Please use DigMontage instead.
warnings.warn(msg, category=DeprecationWarning)
Writing '/home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg/sub-01_ses-01_task-audiovisual_run-01_meg.json'...
{
"TaskName": "audiovisual",
"Manufacturer": "Elekta",
"PowerLineFrequency": 50,
"SamplingFrequency": 600.614990234375,
"SoftwareFilters": "n/a",
"RecordingDuration": 277.7136813300495,
"RecordingType": "continuous",
"DewarPosition": "n/a",
"DigitizedLandmarks": false,
"DigitizedHeadPoints": false,
"MEGChannelCount": 306,
"MEGREFChannelCount": 0,
"EEGChannelCount": 60,
"EOGChannelCount": 1,
"ECGChannelCount": 0,
"EMGChannelCount": 0,
"MiscChannelCount": 0,
"TriggerChannelCount": 9
}
Writing '/home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg/sub-01_ses-01_task-audiovisual_run-01_channels.tsv'...
name type units low_cutoff high_cutoff description sampling_frequency status
MEG 0113 MEGGRADPLANAR T/m 0.10000000149011612 172.17630004882812 Planar Gradiometer 600.614990234375 good
MEG 0112 MEGGRADPLANAR T/m 0.10000000149011612 172.17630004882812 Planar Gradiometer 600.614990234375 good
MEG 0111 MEGMAG T 0.10000000149011612 172.17630004882812 Magnetometer 600.614990234375 good
MEG 0122 MEGGRADPLANAR T/m 0.10000000149011612 172.17630004882812 Planar Gradiometer 600.614990234375 good
MEG 0123 MEGGRADPLANAR T/m 0.10000000149011612 172.17630004882812 Planar Gradiometer 600.614990234375 good
Copying data files to /home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg/sub-01_ses-01_task-audiovisual_run-01_meg.fif
Writing /home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg/sub-01_ses-01_task-audiovisual_run-01_meg.fif
Closing /home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg/sub-01_ses-01_task-audiovisual_run-01_meg.fif [done]
|MNE-sample-data-bids/
|--- participants.tsv
|--- dataset_description.json
|--- participants.json
|--- sub-01/
|------ ses-01/
|--------- sub-01_ses-01_scans.tsv
|--------- meg/
|------------ sub-01_ses-01_coordsystem.json
|------------ sub-01_ses-01_task-audiovisual_run-01_meg.json
|------------ sub-01_ses-01_task-audiovisual_run-01_channels.tsv
|------------ sub-01_ses-01_task-audiovisual_run-01_meg.fif
|------------ sub-01_ses-01_task-audiovisual_run-01_events.tsv
Now let’s assume that we have also collected some T1 weighted MRI data for
our subject. And furthermore, that we have already aligned our coordinate
frames (using e.g., the coregistration GUI) and obtained a transformation
matrix trans.
# Get the path to our MRI scan
t1_mgh_fname = op.join(data_path, 'subjects', 'sample', 'mri', 'T1.mgz')
# Load the transformation matrix and show what it looks like
trans_fname = op.join(data_path, 'MEG', 'sample',
'sample_audvis_raw-trans.fif')
trans = mne.read_trans(trans_fname)
print(trans)
Out:
<Transform | head->MRI (surface RAS)>
[[ 0.99930954 0.01275934 0.0348942 0.00206991]
[ 0.00998479 0.81240475 -0.58300853 0.01130214]
[-0.03578702 0.58295429 0.81171638 -0.02755522]
[ 0. 0. 0. 1. ]]
We can save the MRI to our existing BIDS directory and at the same time
create a JSON sidecar file that contains metadata, we will later use to
retrieve our transformation matrix trans.
# We use the write_anat function
anat_dir = write_anat(bids_root=output_path, # the BIDS dir we wrote earlier
subject=sub,
t1w=t1_mgh_fname, # path to the MRI scan
session=ses,
raw=raw, # the raw MEG data file connected to the MRI
trans=trans, # our transformation matrix
verbose=True # this will print out the sidecar file
)
# Let's have another look at our BIDS directory
print_dir_tree(output_path)
Out:
Writing '/home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/anat/sub-01_ses-01_T1w.json'...
{
"AnatomicalLandmarkCoordinates": {
"LPA": [
197.25741411263368,
153.0008593581418,
138.5894600936019
],
"NAS": [
124.62090614299716,
95.74083565348268,
222.65942693440599
],
"RPA": [
50.71437932937833,
158.24882153365422,
140.05367187187042
]
}
}
|MNE-sample-data-bids/
|--- participants.tsv
|--- dataset_description.json
|--- participants.json
|--- sub-01/
|------ ses-01/
|--------- sub-01_ses-01_scans.tsv
|--------- meg/
|------------ sub-01_ses-01_coordsystem.json
|------------ sub-01_ses-01_task-audiovisual_run-01_meg.json
|------------ sub-01_ses-01_task-audiovisual_run-01_channels.tsv
|------------ sub-01_ses-01_task-audiovisual_run-01_meg.fif
|------------ sub-01_ses-01_task-audiovisual_run-01_events.tsv
|--------- anat/
|------------ sub-01_ses-01_T1w.nii.gz
|------------ sub-01_ses-01_T1w.json
Our BIDS dataset is now ready to be shared. We can easily estimate the
transformation matrix using MNE-BIDS and the BIDS dataset.
bids_fname = bids_basename + '_meg.fif'
# reproduce our trans
estim_trans = get_head_mri_trans(bids_fname=bids_fname, # name of the MEG file
bids_root=output_path # root of our BIDS dir
)
Out:
Opening raw data file /home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg/sub-01_ses-01_task-audiovisual_run-01_meg.fif...
Read a total of 3 projection items:
PCA-v1 (1 x 102) idle
PCA-v2 (1 x 102) idle
PCA-v3 (1 x 102) idle
Range : 25800 ... 192599 = 42.956 ... 320.670 secs
Ready.
Current compensation grade : 0
Reading events from /home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg/sub-01_ses-01_task-audiovisual_run-01_events.tsv.
Reading channel info from /home/stefanappelhoff/mne_data/MNE-sample-data-bids/sub-01/ses-01/meg/sub-01_ses-01_task-audiovisual_run-01_channels.tsv.
/home/stefanappelhoff/Desktop/bids/mne-bids/mne_bids/read.py:165: RuntimeWarning: The unit for channel(s) STI 001, STI 002, STI 003, STI 004, STI 005, STI 006, STI 014, STI 015, STI 016 has changed from V to NA.
raw.set_channel_types(channel_type_dict)
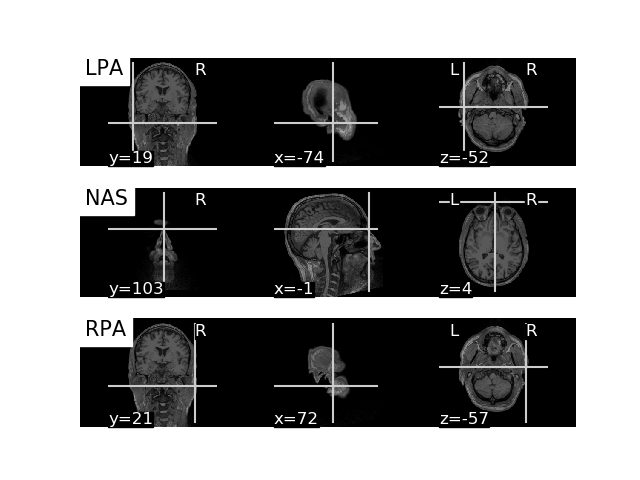
Finally, let’s use the T1 weighted MRI image and plot the anatomical
landmarks Nasion, LPA, and RPA (=left and right preauricular points) onto
the brain image. For that, we can extract the location of Nasion, LPA, and
RPA from the MEG file, apply our transformation matrix trans, and
plot the results.
# Get Landmarks from MEG file, 0, 1, and 2 correspond to LPA, NAS, RPA
# and the 'r' key will provide us with the xyz coordinates
pos = np.asarray((raw.info['dig'][0]['r'],
raw.info['dig'][1]['r'],
raw.info['dig'][2]['r']))
# We use a function from MNE-Python to convert MEG coordinates to MRI space
# for the conversion we use our estimated transformation matrix and the
# MEG coordinates extracted from the raw file. `subjects` and `subjects_dir`
# are used internally, to point to the T1-weighted MRI file: `t1_mgh_fname`
mri_pos = head_to_mri(pos=pos,
subject='sample',
mri_head_t=estim_trans,
subjects_dir=op.join(data_path, 'subjects')
)
# Our MRI written to BIDS, we got `anat_dir` from our `write_anat` function
t1_nii_fname = op.join(anat_dir, 'sub-01_ses-01_T1w.nii.gz')
# Plot it
fig, axs = plt.subplots(3, 1)
for point_idx, label in enumerate(('LPA', 'NAS', 'RPA')):
plot_anat(t1_nii_fname, axes=axs[point_idx],
cut_coords=mri_pos[point_idx, :],
title=label)
plt.show()
Out:
/home/stefanappelhoff/Desktop/bids/mne-bids/examples/convert_mri_and_trans.py:156: UserWarning: Matplotlib is currently using agg, which is a non-GUI backend, so cannot show the figure.
plt.show()
Total running time of the script: ( 0 minutes 3.876 seconds)