mne_connectivity.vector_auto_regression#
- mne_connectivity.vector_auto_regression(data, times=None, names=None, lags=1, l2_reg=0.0, compute_fb_operator=False, model='dynamic', n_jobs=1, verbose=None)[source]#
Compute vector auto-regresssive (VAR) model.
- Parameters:
- dataarray_like, shape (n_epochs, n_signals, n_times) |
Epochs| generator The data from which to compute connectivity. The epochs dimension is interpreted differently, depending on the
'model'argument.- timesarray_like |
None The time points used to construct the epoched
data. IfNone, thentimes_usedin the returnedconnwill not be available.- namesarray_like |
None The names of the nodes of the dataset used to compute connectivity. If
None(default), then names will be a list of integers from 0 ton_nodes. If a list of names, then it must be equal in length ton_nodes.- lags
int|None Autoregressive model order (default 1).
- l2_reg
float|None Ridge penalty (l2-regularization) parameter (default 0.0).
- compute_fb_operatorbool
Whether to compute the backwards operator and average with the forward operator. Addresses bias in the least-square estimation [1].
- model
'dynamic'|'avg-epochs' Whether to compute one VAR model using all epochs as multiple samples of the same VAR model (
'avg-epochs'), or to compute a separate VAR model for each epoch ('dynamic', default), which results in a time-varying VAR model. See Notes.- n_jobs
int The number of jobs to run in parallel (default 1). Requires the joblib package.
- verbosebool |
str|int|None If not
None, override default verbose level (seemne.verbose()for more info). If used, it should be passed as a keyword-argument only.
- dataarray_like, shape (n_epochs, n_signals, n_times) |
- Returns:
- conn
Connectivity|TemporalConnectivity|EpochConnectivity|EpochTemporalConnectivity The connectivity data estimated.
- conn
See also
Notes
Names can be passed in, which are then used to instantiate the nodes of the connectivity class. For example, they can be the electrode names of EEG.
For higher-order VAR models, there are
n_orderAmatrices, representing the linear dynamics with respect to that lag. These are represented by vertically concatenated matrices. For example, if the input is data wheren_signalsis 3, then an order-1 VAR model will result in a 3x3 connectivity matrix. An order-2 VAR model will result in a 6x3 connectivity matrix, with two 3x3 matrices representing the dynamics at lag 1 and lag 2, respectively.When computing a VAR model (i.e. linear dynamical system), we require the input to be a
(n_epochs, n_signals, n_times)3D array. There are two ways one can interpret the data in the model.First, epochs can be treated as multiple samples observed for a single VAR model. That is, we have \(X_1, X_2, ..., X_n\), where each \(X_i\) is a
(n_signals, n_times)data array, withn_epochs. We are interested in estimating the parameters, \((A_1, A_2, ..., A_{order})\), from the following model over all epochs:\[X(t+1) = \sum_{i=0}^{order} A_i X(t-i)\]This results in one VAR model over all the epochs.
The second approach treats each epoch as a different VAR model, estimating a time-varying VAR model. Using the same data as above, we now are interested in estimating the parameters, \((A_1, A_2, ..., A_{order})\) for each epoch. The model would be the following for each epoch:
\[X(t+1) = \sum_{i=0}^{order} A_i X(t-i)\]This results in one VAR model for each epoch. This is done according to the model in [2].
b is of shape [m, m*p], with sub matrices arranged as follows:
b_00
b_01
…
b_0m
b_10
b_11
…
b_1m
…
…
…
…
b_m0
b_m1
…
b_mm
Each sub matrix b_ij is a column vector of length p that contains the filter coefficients from channel j (source) to channel i (sink).
In order to optimize RAM usage, the estimating equations are set up by iterating over sample points. This assumes that there are in general more sample points than channels. You should not estimate a VAR model using less sample points than channels, unless you have good reason.
References
Examples using mne_connectivity.vector_auto_regression#
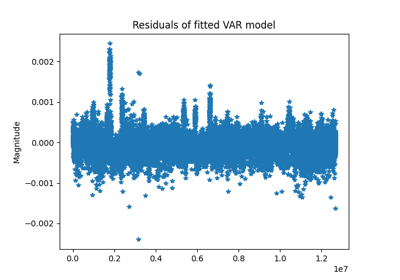
Compute vector autoregressive model (linear system)