Note
Go to the end to download the full example code
Generate simulated evoked data#
Use simulate_sparse_stc() to simulate evoked data.
# Author: Daniel Strohmeier <daniel.strohmeier@tu-ilmenau.de>
# Alexandre Gramfort <alexandre.gramfort@inria.fr>
#
# License: BSD-3-Clause
import numpy as np
import matplotlib.pyplot as plt
import mne
from mne.datasets import sample
from mne.time_frequency import fit_iir_model_raw
from mne.viz import plot_sparse_source_estimates
from mne.simulation import simulate_sparse_stc, simulate_evoked
print(__doc__)
Load real data as templates:
data_path = sample.data_path()
meg_path = data_path / 'MEG' / 'sample'
raw = mne.io.read_raw_fif(meg_path / 'sample_audvis_raw.fif')
proj = mne.read_proj(meg_path / 'sample_audvis_ecg-proj.fif')
raw.add_proj(proj)
raw.info['bads'] = ['MEG 2443', 'EEG 053'] # mark bad channels
fwd_fname = meg_path / 'sample_audvis-meg-eeg-oct-6-fwd.fif'
ave_fname = meg_path / 'sample_audvis-no-filter-ave.fif'
cov_fname = meg_path / 'sample_audvis-cov.fif'
fwd = mne.read_forward_solution(fwd_fname)
fwd = mne.pick_types_forward(fwd, meg=True, eeg=True, exclude=raw.info['bads'])
cov = mne.read_cov(cov_fname)
info = mne.io.read_info(ave_fname)
label_names = ['Aud-lh', 'Aud-rh']
labels = [mne.read_label(meg_path / 'labels' / f'{ln}.label')
for ln in label_names]
Opening raw data file /home/circleci/mne_data/MNE-sample-data/MEG/sample/sample_audvis_raw.fif...
Read a total of 3 projection items:
PCA-v1 (1 x 102) idle
PCA-v2 (1 x 102) idle
PCA-v3 (1 x 102) idle
Range : 25800 ... 192599 = 42.956 ... 320.670 secs
Ready.
Read a total of 6 projection items:
ECG-planar-999--0.200-0.400-PCA-01 (1 x 203) idle
ECG-planar-999--0.200-0.400-PCA-02 (1 x 203) idle
ECG-axial-999--0.200-0.400-PCA-01 (1 x 102) idle
ECG-axial-999--0.200-0.400-PCA-02 (1 x 102) idle
ECG-eeg-999--0.200-0.400-PCA-01 (1 x 59) idle
ECG-eeg-999--0.200-0.400-PCA-02 (1 x 59) idle
6 projection items deactivated
Reading forward solution from /home/circleci/mne_data/MNE-sample-data/MEG/sample/sample_audvis-meg-eeg-oct-6-fwd.fif...
Reading a source space...
Computing patch statistics...
Patch information added...
Distance information added...
[done]
Reading a source space...
Computing patch statistics...
Patch information added...
Distance information added...
[done]
2 source spaces read
Desired named matrix (kind = 3523) not available
Read MEG forward solution (7498 sources, 306 channels, free orientations)
Desired named matrix (kind = 3523) not available
Read EEG forward solution (7498 sources, 60 channels, free orientations)
Forward solutions combined: MEG, EEG
Source spaces transformed to the forward solution coordinate frame
364 out of 366 channels remain after picking
366 x 366 full covariance (kind = 1) found.
Read a total of 4 projection items:
PCA-v1 (1 x 102) active
PCA-v2 (1 x 102) active
PCA-v3 (1 x 102) active
Average EEG reference (1 x 60) active
Read a total of 4 projection items:
PCA-v1 (1 x 102) active
PCA-v2 (1 x 102) active
PCA-v3 (1 x 102) active
Average EEG reference (1 x 60) active
Generate source time courses from 2 dipoles and the correspond evoked data
times = np.arange(300, dtype=np.float64) / raw.info['sfreq'] - 0.1
rng = np.random.RandomState(42)
def data_fun(times):
"""Function to generate random source time courses"""
return (50e-9 * np.sin(30. * times) *
np.exp(- (times - 0.15 + 0.05 * rng.randn(1)) ** 2 / 0.01))
stc = simulate_sparse_stc(fwd['src'], n_dipoles=2, times=times,
random_state=42, labels=labels, data_fun=data_fun)
Generate noisy evoked data
picks = mne.pick_types(raw.info, meg=True, exclude='bads')
iir_filter = fit_iir_model_raw(raw, order=5, picks=picks, tmin=60, tmax=180)[1]
nave = 100 # simulate average of 100 epochs
evoked = simulate_evoked(fwd, stc, info, cov, nave=nave, use_cps=True,
iir_filter=iir_filter)
Average patch normals will be employed in the rotation to the local surface coordinates....
Converting to surface-based source orientations...
[done]
Projecting source estimate to sensor space...
[done]
4 projection items deactivated
Created an SSP operator (subspace dimension = 4)
4 projection items activated
SSP projectors applied...
Plot
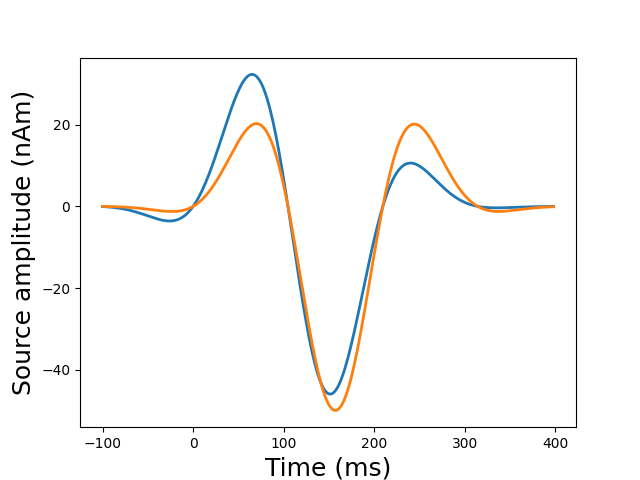
plot_sparse_source_estimates(fwd['src'], stc, bgcolor=(1, 1, 1),
opacity=0.5, high_resolution=True)
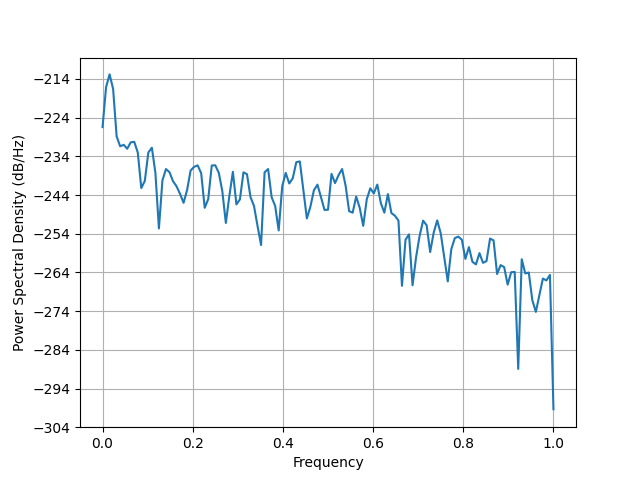
plt.figure()
plt.psd(evoked.data[0])
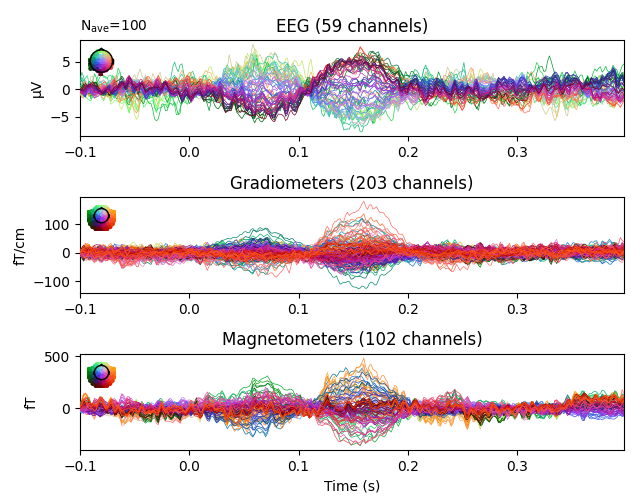
evoked.plot(time_unit='s')
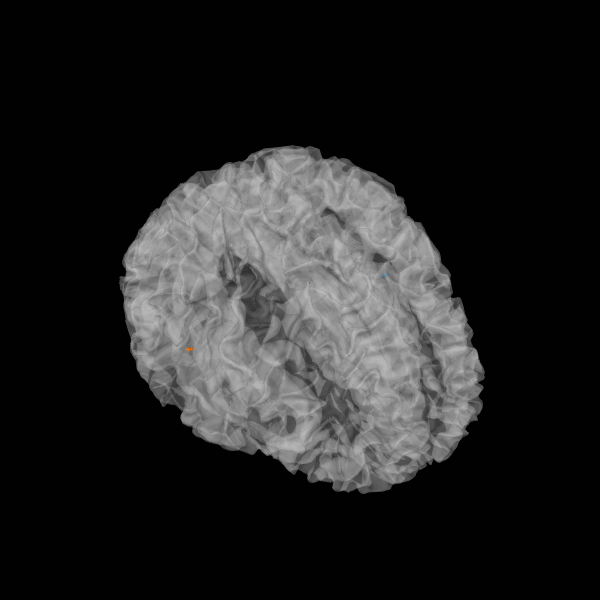
Total number of active sources: 2
Total running time of the script: ( 0 minutes 7.646 seconds)
Estimated memory usage: 295 MB