mne.pick_types_forward#
- mne.pick_types_forward(orig, meg=False, eeg=False, ref_meg=True, seeg=False, ecog=False, dbs=False, include=[], exclude=[])[source]#
Pick by channel type and names from a forward operator.
- Parameters:
- orig
dict A forward solution.
- meg
bool|str If True include MEG channels. If string it can be ‘mag’, ‘grad’, ‘planar1’ or ‘planar2’ to select only magnetometers, all gradiometers, or a specific type of gradiometer.
- eeg
bool If True include EEG channels.
- ref_meg
bool If True include CTF / 4D reference channels.
- seeg
bool If True include stereotactic EEG channels.
- ecog
bool If True include electrocorticography channels.
- dbs
bool If True include deep brain stimulation channels.
- include
listofstr List of additional channels to include. If empty do not include any.
- exclude
listofstr|str List of channels to exclude. If empty do not exclude any (default). If ‘bads’, exclude channels in orig[‘info’][‘bads’].
- orig
- Returns:
- res
dict Forward solution restricted to selected channel types.
- res
Examples using mne.pick_types_forward#
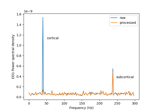
Cortical Signal Suppression (CSS) for removal of cortical signals
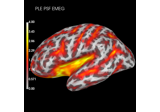
Compute spatial resolution metrics to compare MEG with EEG+MEG