mne_bids.write_raw_bids#
- mne_bids.write_raw_bids(raw, bids_path, events=None, event_id=None, event_metadata=None, extra_columns_descriptions=None, *, anonymize=None, format='auto', physical_range='auto', symlink=False, empty_room=None, allow_preload=False, montage=None, acpc_aligned=False, electrodes_tsv_task=False, emg_placement=None, overwrite=False, readme=True, verbose=None)[source]#
Save raw data to a BIDS-compliant folder structure.
Warning
The original file is simply copied over if the original file format is BIDS-supported for that datatype. Otherwise, this function will convert to a BIDS-supported file format while warning the user. For EEG and iEEG data, conversion will be to BrainVision format; for MEG, conversion will be to FIFF.
mne-bidswill infer the manufacturer information from the file extension. If your file format is non-standard for the manufacturer, please update the manufacturer field in the sidecars manually.
- Parameters:
- raw
mne.io.Raw The raw data. It must be an instance of
mne.io.Rawthat is not already loaded from disk unlessallow_preloadis explicitly set toTrue. See warning for theallow_preloadparameter.- bids_path
BIDSPath The file to write. The
mne_bids.BIDSPathinstance passed here must have thesubject,task, androotattributes set. If thedatatypeattribute is not set, it will be inferred from the recording data type found inraw. In case of multiple data types, the.datatypeattribute must be set. Example:bids_path = BIDSPath(subject='01', session='01', task='testing', acquisition='01', run='01', datatype='meg', root='/data/BIDS')
This will write the following files in the correct subfolder
root:sub-01_ses-01_task-testing_acq-01_run-01_meg.fif sub-01_ses-01_task-testing_acq-01_run-01_meg.json sub-01_ses-01_task-testing_acq-01_run-01_channels.tsv sub-01_ses-01_acq-01_coordsystem.json
and the following one if
eventsis notNone:sub-01_ses-01_task-testing_acq-01_run-01_events.tsv
and add a line to the following files:
participants.tsv scans.tsv
Note that the extension is automatically inferred from the raw object.
- eventspath-like |
np.ndarray|None Use this parameter to specify events to write to the
*_events.tsvsidecar file, additionally to the object’sAnnotations(which are always written). Ifpath-like, specifies the location of an MNE events file. If an array, the MNE events array (shape:(n_events, 3)). If a path or an array andraw.annotationsexist, the union ofeventsandraw.annotationswill be written. Mappings from event names to event codes (listed in the third column of the MNE events array) must be specified via theevent_idparameter; otherwise, an exception is raised. IfAnnotationsare present, their descriptions must be included inevent_idas well. IfNone, events will only be inferred from the raw object’sAnnotations.Note
If specified, writes the union of
eventsandraw.annotations. If you wish to only writeraw.annotations, passevents=None. If you want to exclude the events inraw.annotationsfrom being written, callraw.set_annotations(None)before invoking this function.Note
Either, descriptions of all event codes must be specified via the
event_idparameter or each event must be accompanied by a row inevent_metadata.- event_id
dict|None Descriptions or names describing the event codes, if you passed
events. The descriptions will be written to thetrial_typecolumn in*_events.tsv. The dictionary keys correspond to the event description,s and the values to the event codes. You must specify a description for all event codes appearing inevents. If your data containsAnnotations, you can use this parameter to assign event codes to each unique annotation description (mapping from description to event code).- event_metadata
pandas.DataFrame|None Metadata for each event in
events. Each row corresponds to an event.- extra_columns_descriptions
dict|None A dictionary that maps column names of the
event_metadatato descriptions. Each column ofevent_metadatamust have a corresponding entry in this.- anonymize
dict|None If
None(default), no anonymization is performed. If a dictionary, data will be anonymized depending on the dictionary keys:daysbackis a required key,keep_hisis optional.daysbackintNumber of days by which to move back the recording date in time. In studies with multiple subjects the relative recording date differences between subjects can be kept by using the same number of
daysbackfor all subject anonymizations.daysbackshould be great enough to shift the date prior to 1925 to conform with BIDS anonymization rules.keep_hisboolIf
False(default), all subject information next to the recording date will be overwritten as well. IfTrue, keep subject information apart from the recording date.keep_sourceboolWhether to store the name of the
rawinput file in thesourcecolumn ofscans.tsv. By default, this information is not stored.
- format‘auto’ | ‘BrainVision’ | ‘BDF’ | ‘EDF’ | ‘FIF’ | ‘EEGLAB’
Controls the file format of the data after BIDS conversion. If
'auto', MNE-BIDS will attempt to convert the input data to BIDS without a change of the original file format. A conversion to a different file format will then only take place if the original file format lacks some necessary features. Conversion may be forced to BrainVision, BDF, EDF, or EEGLAB for EEG, to BrainVision, EDF, or EEGLAB for iEEG, to BDF or EDF for EMG, and to FIF for MEG data.- physical_range
str|tuple If
'auto'(default), the physical range is inferred from the data, taking the minimum and maximum values per channel type. If'channelwise', the range will be defined per channel. If a tuple of minimum and maximum, this manual physical range will be used. Only used for exporting EDF files.Added in version 0.19.
- symlink
bool Instead of copying the source files, only create symbolic links to preserve storage space. This is only allowed when not anonymizing the data (i.e.,
anonymizemust beNone).Note
Symlinks currently only work with FIFF files. In case of split files, only a link to the first file will be created, and
mne_bids.read_raw_bids()will correctly handle reading the data again.Note
Symlinks are currently only supported on macOS and Linux. We will add support for Windows 10 at a later time.
- empty_room
mne.io.Raw|BIDSPath|None The empty-room recording to be associated with this file. This is only supported for MEG data. If
Raw, you may pass raw data that was not preloaded (otherwise, passallow_preload=True); i.e., it behaves similar to therawparameter. The session name will be automatically generated from the raw object’sinfo['meas_date']. If aBIDSPath, therootattribute must be the same as inbids_path. PassNone(default) if you do not wish to specify an associated empty-room recording.Changed in version 0.11: Accepts
Rawdata.- allow_preload
bool If
True, allow writing of preloaded raw objects (i.e.,raw.preloadisTrue). Because the original file is ignored, you must specify whatformatto write (notauto).Warning
BIDS was originally designed for unprocessed or minimally processed data. For this reason, by default, we prevent writing of preloaded data that may have been modified. Only use this option when absolutely necessary: for example, manually converting from file formats not supported by MNE or writing preprocessed derivatives. Be aware that these use cases are not fully supported.
- montage
mne.channels.DigMontage|None The montage with channel positions if channel position data are to be stored in a format other than “head” (the internal MNE coordinate frame that the data in
rawis stored in).- acpc_aligned
bool It is difficult to check whether the T1 scan is ACPC aligned which means that “mri” coordinate space is “ACPC” BIDS coordinate space. So, this flag is required to be True when the digitization data is in “mri” for intracranial data to confirm that the T1 is ACPC-aligned.
- electrodes_tsv_task
bool Add the
task-entity to theelectrodes.tsvfilename. Defaults toFalse.- emg_placement“Measured” | “ChannelSpecific” | “Other” |
None How the EMG sensor locations were determined. Must be one of the literal strings if datatype is “emg” and should be
Nonefor all other datatypes.- overwrite
bool Whether to overwrite existing files or data in files. Defaults to
False.If
True, any existing files with the same BIDS parameters will be overwritten with the exception of the*_participants.tsvand*_scans.tsvfiles. For these files, parts of pre-existing data that match the current data will be replaced. For*_participants.tsv, specifically, age, sex and hand fields will be overwritten, while any manually added fields inparticipants.jsonandparticipants.tsvby a user will be retained. IfFalse, no existing data will be overwritten or replaced.- readme
bool If
False, leave any existingREADMEuntouched and do not create one. Defaults toTrue.Added in version 0.19.
- verbose
bool|str|int|None Control verbosity of the logging output. If
None, use the default verbosity level. See the logging documentation andmne.verbose()for details. Should only be passed as a keyword argument.
- raw
- Returns:
Notes
You should ensure that
raw.info['subject_info']andraw.info['meas_date']are set to proper (not-None) values to allow for the correct computation of each participant’s age when creating*_participants.tsv.This function will convert existing
mne.Annotationsfromraw.annotationsto events. Additionally, any events supplied viaeventswill be written too. To avoid writing of annotations, remove them from the raw file viaraw.set_annotations(None)before invokingwrite_raw_bids.To write events encoded in a
STIMchannel, you first need to create the events array manually and pass it to this function:See the documentation of
mne.find_events()for more information on event extraction fromSTIMchannels.When anonymizing
.edffiles, then the file format for EDF limits how far back we can set the recording date. Therefore, all anonymized EDF datasets will have an internal recording date of01-01-1985, and the actual recording date will be stored in thescans.tsvfile’sacq_timecolumn.write_raw_bidswill generate adataset_description.jsonfile if it does not already exist. Minimal metadata will be written there. If one setsoverwritetoTruehere, it will not overwrite an existingdataset_description.jsonfile. If you need to add more data there, or overwrite it, then you should callmne_bids.make_dataset_description()directly.When writing EDF or BDF files, all file extensions are forced to be lower-case, in compliance with the BIDS specification.
Examples using mne_bids.write_raw_bids#
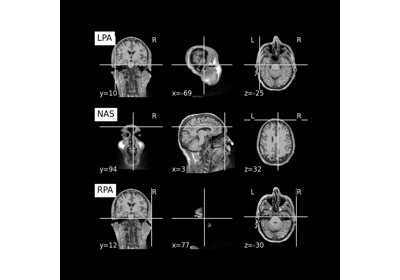
Save and load T1-weighted MRI scan along with anatomical landmarks in BIDS
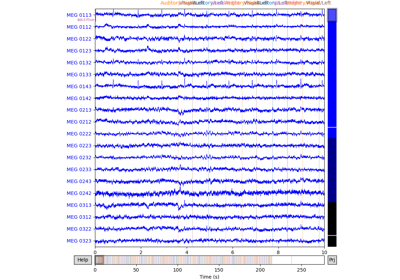
Interactive data inspection and bad channel selection