mne_nirs.statistics.RegressionResults#
- class mne_nirs.statistics.RegressionResults(info, data, design)[source]#
Class containing GLM regression results.
- Parameters:
- Attributes:
ch_namesReturn the channel names.
compensation_gradeThe current gradient compensation grade.
- data
Methods
MSE()Return the GLM MSE.
compute_contrast(contrast[, contrast_type])Compute contrasts on regression results.
copy()Return a copy of the GLM results.
get_channel_types([picks, unique, only_data_chs])Get a list of channel type for each channel.
model()Return the GLM model.
pick(picks[, exclude])Pick a subset of channels.
plot_topo([conditions, axes, vlim, vmin, ...])Plot 2D topography of GLM data.
save(fname[, overwrite])Save GLM results to disk.
scatter([conditions, exclude_no_interest, ...])Scatter plot of the GLM results.
surface_projection([chroma, condition, ...])Project GLM results on to the surface of the brain.
theta()Return the GLM theta results.
to_dataframe([order])Return a tidy dataframe representing the GLM results.
to_dataframe_region_of_interest(group_by, ...)Region of interest results as a dataframe.
- Returns:
- glm_estResultsGLM,
Result class.
- MSE()[source]#
Return the GLM MSE.
- Returns:
- thetas
array Array of MSEs. A value is provided per channel.
- thetas
- __contains__(ch_type)#
Check channel type membership.
- Parameters:
- ch_type
str Channel type to check for. Can be e.g.
'meg','eeg','stim', etc.
- ch_type
- Returns:
- inbool
Whether or not the instance contains the given channel type.
Examples
Channel type membership can be tested as:
>>> 'meg' in inst True >>> 'seeg' in inst False
- __hash__ = None#
- property compensation_grade#
The current gradient compensation grade.
- compute_contrast(contrast, contrast_type=None)[source]#
Compute contrasts on regression results.
This is a wrapper function for nilearn.stats.contrasts.
- Parameters:
- contrast
numpy.ndarrayof shape (p) or (q, p), Where q = number of contrast vectors and p = number of regressors.
- contrast_type{
None, ‘t’, ‘F’}, optional Type of the contrast. If None, then defaults to ‘t’ for 1D con_val and ‘F’ for 2D con_val.
- contrast
- Returns:
- contrastContrast instance,
Yields the statistics of the contrast (effects, variance, p-values).
- copy()#
Return a copy of the GLM results.
- Returns:
- instinstance of ResultsGLM
A copy of the object.
- get_channel_types(picks=None, unique=False, only_data_chs=False)#
Get a list of channel type for each channel.
- Parameters:
- picks
str| array_like |slice|None Channels to include. Slices and lists of integers will be interpreted as channel indices. In lists, channel type strings (e.g.,
['meg', 'eeg']) will pick channels of those types, channel name strings (e.g.,['MEG0111', 'MEG2623']will pick the given channels. Can also be the string values'all'to pick all channels, or'data'to pick data channels. None (default) will pick all channels. Note that channels ininfo['bads']will be included if their names or indices are explicitly provided.- uniquebool
Whether to return only unique channel types. Default is
False.- only_data_chsbool
Whether to ignore non-data channels. Default is
False.
- picks
- Returns:
- channel_types
list The channel types.
- channel_types
- model()[source]#
Return the GLM model.
- Returns:
- models
array Array of models. An model is provided per channel.
- models
- pick(picks, exclude=())[source]#
Pick a subset of channels.
- Parameters:
- picks
str| array_like |slice|None Channels to include. Slices and lists of integers will be interpreted as channel indices. In lists, channel type strings (e.g.,
['meg', 'eeg']) will pick channels of those types, channel name strings (e.g.,['MEG0111', 'MEG2623']will pick the given channels. Can also be the string values'all'to pick all channels, or'data'to pick data channels. None (default) will pick all channels. Note that channels ininfo['bads']will be included if their names or indices are explicitly provided.- exclude
list|str Set of channels to exclude, only used when picking based on types (e.g., exclude=”bads” when picks=”meg”).
- picks
- Returns:
- instinstance of ResultsGLM
The modified instance.
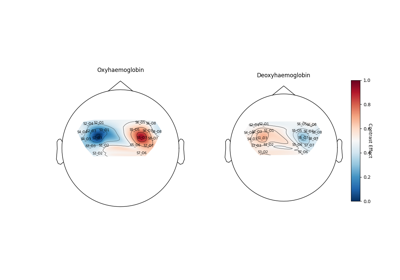
- plot_topo(conditions=None, axes=None, *, vlim=(None, None), vmin=None, vmax=None, colorbar=True, figsize=(12, 7), sphere=None)[source]#
Plot 2D topography of GLM data.
- Parameters:
- conditions
array Which conditions should be displayed.
- axesinstance of Axes |
None The axes to plot to. If None, a new figure is used.
- vlim
tupleof length 2 Colormap limits to use. If a
tupleof floats, specifies the lower and upper bounds of the colormap (in that order); providingNonefor either entry will set the corresponding boundary at the positive or negative max of the absolute value of the data.- vmin
float|None Deprecated, use vlim instead.
- vmax
float|None Deprecated, use vlim instead.
- colorbarBool
Should a colorbar be plotted.
- figsizetwo values
Figure size.
- sphereAs specified in MNE
Sphere parameter from mne.viz.topomap.plot_topomap.
- conditions
- Returns:
- figfigure
Figure of each design matrix component for hbo (top row) and hbr (bottom row).
- save(fname, overwrite=False)#
Save GLM results to disk.
- scatter(conditions=(), exclude_no_interest=True, axes=None, no_interest=None)#
Scatter plot of the GLM results.
- Parameters:
- conditions
list List of condition names to plot. By default plots all regressors of interest.
- exclude_no_interestbool
Exclude regressors of no interest from the figure.
- axesAxes
Optional axes on which to plot the data.
- no_interest
list List of regressors that are of no interest. If none are specified then conditions starting with [“drift”, “constant”, “short”, “Short”] will be excluded.
- conditions
- Returns:
- pltmatplotlib.Figure
Scatter plot.
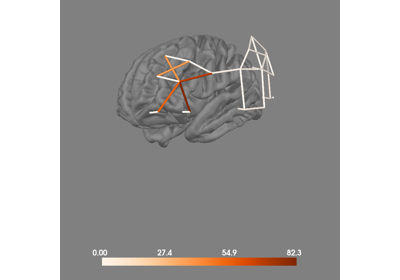
- surface_projection(chroma='hbo', condition=None, background='w', figure=None, clim='auto', mode='weighted', colormap='RdBu_r', surface='pial', hemi='both', size=800, view=None, colorbar=True, distance=0.03, subjects_dir=None, src=None, verbose=False)[source]#
Project GLM results on to the surface of the brain.
Note: This function provides a convenient wrapper around low level MNE-Python functions. It is convenient if you wish to use a generic head model. If you have acquired fMRI images you may wish to use the underlying lower level functions.
Note: This function does not conduct a forward model analysis with photon migration etc. It simply projects the values from each channel to the local cortical surface. It is useful for visualisation, but users should take this in to consideration when drawing conclusions from the visualisation.
- Parameters:
- chroma
str Chromophore to plot`.
- condition
str Condition to plot`.
- background
matplotlibcolor Color of the background of the display window.
- figuremayavi.core.api.Scene,
matplotlib.figure.Figure,list,None If None, a new figure will be created. If multiple views or a split view is requested, this must be a list of the appropriate length. If int is provided it will be used to identify the Mayavi figure by it’s id or create a new figure with the given id. If an instance of matplotlib figure, mpl backend is used for plotting.
- clim
str|dict Colorbar properties specification. If ‘auto’, set clim automatically based on data percentiles. If dict, should contain:
kind‘value’ | ‘percent’Flag to specify type of limits.
limslist | np.ndarray | tuple of float, 3 elementsLower, middle, and upper bounds for colormap.
pos_limslist | np.ndarray | tuple of float, 3 elementsLower, middle, and upper bound for colormap. Positive values will be mirrored directly across zero during colormap construction to obtain negative control points.
Note
Only one of
limsorpos_limsshould be provided. Only sequential colormaps should be used withlims, and only divergent colormaps should be used withpos_lims.- mode
str Can be “sum” to do a linear sum of weights, “weighted” to make this a weighted sum, “nearest” to use only the weight of the nearest sensor, or “single” to do a distance-weight of the nearest sensor. Default is “sum”.
- colormap
str Colormap to use.
- surface
str The type of surface (inflated, white etc.).
- hemi
str Hemisphere id (ie ‘lh’, ‘rh’, ‘both’, or ‘split’). In the case of ‘both’, both hemispheres are shown in the same window. In the case of ‘split’ hemispheres are displayed side-by-side in different viewing panes.
- size
floatortupleoffloat The size of the window, in pixels. can be one number to specify a square window, or the (width, height) of a rectangular window. Has no effect with mpl backend.
- view
str View to set brain to.
- colorbarbool
If True, display colorbar on scene.
- distance
float Distance (m) defining the activation “ball” of the sensor.
- subjects_dirpath-like |
None The path to the directory containing the FreeSurfer subjects reconstructions. If
None, defaults to theSUBJECTS_DIRenvironment variable.- srcinstance of
SourceSpaces The source space.
- verbosebool |
str|int|None Control verbosity of the logging output. If
None, use the default verbosity level. See the logging documentation andmne.verbose()for details. Should only be passed as a keyword argument.
- chroma
- Returns:
- figureinstance of
mne.viz.Brain|matplotlib.figure.Figure An instance of
mne.viz.Brainor matplotlib figure.
- figureinstance of
- theta()[source]#
Return the GLM theta results.
- Returns:
- thetas
array Array of thetas. An array is provided per channel.
- thetas
- to_dataframe(order=None)#
Return a tidy dataframe representing the GLM results.
- Parameters:
- order
list Order in which the rows should be returned by channel name.
- order
- Returns:
- tidy
pandas.DataFrame Dataframe containing GLM results.
- tidy
- to_dataframe_region_of_interest(group_by, condition, weighted=True, demographic_info=False)[source]#
Region of interest results as a dataframe.
- Parameters:
- group_by
dict Specifies which channels are aggregated into a single ROI. The dict key will be used as the ROI label and the dict values must be lists of picks (either channel names or integer indices of
epochs.ch_names). For example:group_by=dict(Left_ROI=[1, 2, 3, 4], Right_ROI=[5, 6, 7, 8])
- condition
str|list Name to be used for condition.
- weightedBool |
dict Weighting to be applied to each channel in the ROI computation. If False, then all channels will be weighted equally. If True, channels will be weighted by the inverse of the standard error of the GLM fit. For manual specification of the channel weighting a dictionary can be provided. If a dictionary is provided, the keys and length of lists must match the
group_byparameters. The weights will be scaled internally to sum to 1.- demographic_infoBool
Add an extra column with demographic information from info[“subject_info”].
- group_by
- Returns:
- tidy
pandas.DataFrame Dataframe containing GLM results.
- tidy