mne_nirs.statistics.run_glm#
- mne_nirs.statistics.run_glm(raw, design_matrix, noise_model='ar1', bins=0, n_jobs=1, verbose=0)[source]#
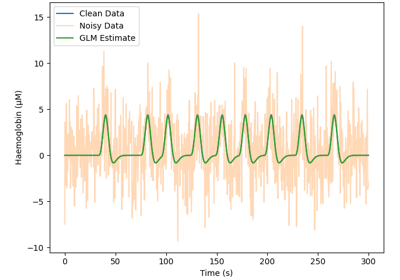
GLM fit for an MNE structure containing fNIRS data.
This is a wrapper function for nilearn.stats.first_level_model.run_glm.
- Parameters:
- rawinstance of
Raw The haemoglobin data.
- design_matrixas specified in Nilearn
The design matrix as generated by mne_nirs.make_first_level_design_matrix. See example
9.5.5. Examples of design matricesat https://nilearn.github.io/auto_examples/index.html for details on how to specify design matrices.- noise_model{‘ar1’, ‘ols’, ‘arN’, ‘auto’}, optional
The temporal variance model. Defaults to first order auto regressive model ‘ar1’. The AR model can be set to any integer value by modifying the value of N. E.g. use ar5 for a fifth order model. If the string auto is provided a model with order 4 times the sample rate will be used.
- bins
int, optional Maximum number of discrete bins for the AR coef histogram/clustering. By default the value is 0, which will set the number of bins to the number of channels, effectively estimating the AR model for each channel.
- n_jobs
int, optional The number of CPUs to use to do the computation. -1 means ‘all CPUs’.
- verbose
int, optional The verbosity level. Default is 0.
- rawinstance of
- Returns:
- glm_estimates
RegressionResults RegressionResults class which stores the GLM results.
- glm_estimates