Note
Click here to download the full example code
Compute sLORETA inverse solution on raw data¶
Compute sLORETA inverse solution on raw dataset restricted to a brain label and stores the solution in stc files for visualisation.
# Author: Alexandre Gramfort <alexandre.gramfort@inria.fr>
#
# License: BSD-3-Clause
import matplotlib.pyplot as plt
import mne
from mne.datasets import sample
from mne.minimum_norm import apply_inverse_raw, read_inverse_operator
print(__doc__)
data_path = sample.data_path()
fname_inv = data_path + '/MEG/sample/sample_audvis-meg-oct-6-meg-inv.fif'
fname_raw = data_path + '/MEG/sample/sample_audvis_raw.fif'
label_name = 'Aud-lh'
fname_label = data_path + '/MEG/sample/labels/%s.label' % label_name
snr = 1.0 # use smaller SNR for raw data
lambda2 = 1.0 / snr ** 2
method = "sLORETA" # use sLORETA method (could also be MNE or dSPM)
# Load data
raw = mne.io.read_raw_fif(fname_raw)
inverse_operator = read_inverse_operator(fname_inv)
label = mne.read_label(fname_label)
raw.set_eeg_reference('average', projection=True) # set average reference.
start, stop = raw.time_as_index([0, 15]) # read the first 15s of data
# Compute inverse solution
stc = apply_inverse_raw(raw, inverse_operator, lambda2, method, label,
start, stop, pick_ori=None)
# Save result in stc files
stc.save('mne_%s_raw_inverse_%s' % (method, label_name))
Out:
Opening raw data file /home/circleci/mne_data/MNE-sample-data/MEG/sample/sample_audvis_raw.fif...
Read a total of 3 projection items:
PCA-v1 (1 x 102) idle
PCA-v2 (1 x 102) idle
PCA-v3 (1 x 102) idle
Range : 25800 ... 192599 = 42.956 ... 320.670 secs
Ready.
Reading inverse operator decomposition from /home/circleci/mne_data/MNE-sample-data/MEG/sample/sample_audvis-meg-oct-6-meg-inv.fif...
Reading inverse operator info...
[done]
Reading inverse operator decomposition...
[done]
305 x 305 full covariance (kind = 1) found.
Read a total of 4 projection items:
PCA-v1 (1 x 102) active
PCA-v2 (1 x 102) active
PCA-v3 (1 x 102) active
Average EEG reference (1 x 60) active
Noise covariance matrix read.
22494 x 22494 diagonal covariance (kind = 2) found.
Source covariance matrix read.
22494 x 22494 diagonal covariance (kind = 6) found.
Orientation priors read.
22494 x 22494 diagonal covariance (kind = 5) found.
Depth priors read.
Did not find the desired covariance matrix (kind = 3)
Reading a source space...
Computing patch statistics...
Patch information added...
Distance information added...
[done]
Reading a source space...
Computing patch statistics...
Patch information added...
Distance information added...
[done]
2 source spaces read
Read a total of 4 projection items:
PCA-v1 (1 x 102) active
PCA-v2 (1 x 102) active
PCA-v3 (1 x 102) active
Average EEG reference (1 x 60) active
Source spaces transformed to the inverse solution coordinate frame
Adding average EEG reference projection.
1 projection items deactivated
Preparing the inverse operator for use...
Scaled noise and source covariance from nave = 1 to nave = 1
Created the regularized inverter
Created an SSP operator (subspace dimension = 3)
Created the whitener using a noise covariance matrix with rank 302 (3 small eigenvalues omitted)
Computing noise-normalization factors (sLORETA)...
[done]
Applying inverse to raw...
Picked 305 channels from the data
Computing inverse...
Eigenleads need to be weighted ...
combining the current components...
[done]
Writing STC to disk...
[done]
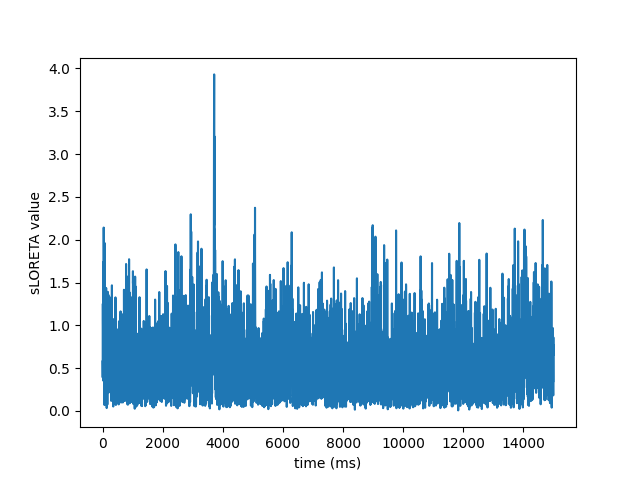
View activation time-series
plt.plot(1e3 * stc.times, stc.data[::100, :].T)
plt.xlabel('time (ms)')
plt.ylabel('%s value' % method)
plt.show()
Total running time of the script: ( 0 minutes 4.647 seconds)
Estimated memory usage: 12 MB