Note
Click here to download the full example code
Permutation F-test on sensor data with 1D cluster level#
One tests if the evoked response is significantly different between conditions. Multiple comparison problem is addressed with cluster level permutation test.
# Authors: Alexandre Gramfort <alexandre.gramfort@inria.fr>
#
# License: BSD-3-Clause
import matplotlib.pyplot as plt
import mne
from mne import io
from mne.stats import permutation_cluster_test
from mne.datasets import sample
print(__doc__)
Set parameters
data_path = sample.data_path()
meg_path = data_path / 'MEG' / 'sample'
raw_fname = meg_path / 'sample_audvis_filt-0-40_raw.fif'
event_fname = meg_path / 'sample_audvis_filt-0-40_raw-eve.fif'
tmin = -0.2
tmax = 0.5
# Setup for reading the raw data
raw = io.read_raw_fif(raw_fname)
events = mne.read_events(event_fname)
channel = 'MEG 1332' # include only this channel in analysis
include = [channel]
Opening raw data file /home/circleci/mne_data/MNE-sample-data/MEG/sample/sample_audvis_filt-0-40_raw.fif...
Read a total of 4 projection items:
PCA-v1 (1 x 102) idle
PCA-v2 (1 x 102) idle
PCA-v3 (1 x 102) idle
Average EEG reference (1 x 60) idle
Range : 6450 ... 48149 = 42.956 ... 320.665 secs
Ready.
Read epochs for the channel of interest
picks = mne.pick_types(raw.info, meg=False, eog=True, include=include,
exclude='bads')
event_id = 1
reject = dict(grad=4000e-13, eog=150e-6)
epochs1 = mne.Epochs(raw, events, event_id, tmin, tmax, picks=picks,
baseline=(None, 0), reject=reject)
condition1 = epochs1.get_data() # as 3D matrix
event_id = 2
epochs2 = mne.Epochs(raw, events, event_id, tmin, tmax, picks=picks,
baseline=(None, 0), reject=reject)
condition2 = epochs2.get_data() # as 3D matrix
condition1 = condition1[:, 0, :] # take only one channel to get a 2D array
condition2 = condition2[:, 0, :] # take only one channel to get a 2D array
Not setting metadata
72 matching events found
Setting baseline interval to [-0.19979521315838786, 0.0] sec
Applying baseline correction (mode: mean)
4 projection items activated
Loading data for 72 events and 106 original time points ...
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
16 bad epochs dropped
Not setting metadata
73 matching events found
Setting baseline interval to [-0.19979521315838786, 0.0] sec
Applying baseline correction (mode: mean)
4 projection items activated
Loading data for 73 events and 106 original time points ...
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
Rejecting epoch based on EOG : ['EOG 061']
11 bad epochs dropped
Compute statistic
threshold = 6.0
T_obs, clusters, cluster_p_values, H0 = \
permutation_cluster_test([condition1, condition2], n_permutations=1000,
threshold=threshold, tail=1, n_jobs=1,
out_type='mask')
stat_fun(H1): min=0.000227 max=38.167093
Running initial clustering
Found 4 clusters
Permuting 999 times...
0%| | : 0/999 [00:00<?, ?it/s]
13%|#2 | : 129/999 [00:00<00:00, 3733.70it/s]
27%|##6 | : 269/999 [00:00<00:00, 3926.71it/s]
42%|####1 | : 415/999 [00:00<00:00, 4062.24it/s]
56%|#####6 | : 562/999 [00:00<00:00, 4136.79it/s]
70%|####### | : 704/999 [00:00<00:00, 4149.49it/s]
85%|########5 | : 851/999 [00:00<00:00, 4186.32it/s]
100%|##########| : 999/999 [00:00<00:00, 4248.05it/s]
100%|##########| : 999/999 [00:00<00:00, 4220.30it/s]
Computing cluster p-values
Done.
Plot
times = epochs1.times
fig, (ax, ax2) = plt.subplots(2, 1, figsize=(8, 4))
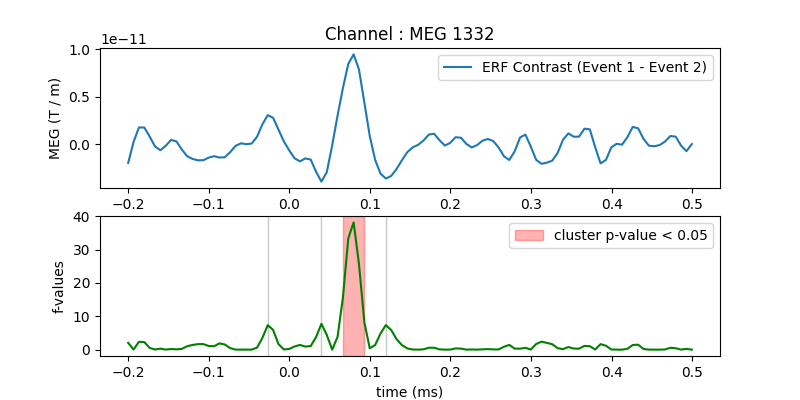
ax.set_title('Channel : ' + channel)
ax.plot(times, condition1.mean(axis=0) - condition2.mean(axis=0),
label="ERF Contrast (Event 1 - Event 2)")
ax.set_ylabel("MEG (T / m)")
ax.legend()
for i_c, c in enumerate(clusters):
c = c[0]
if cluster_p_values[i_c] <= 0.05:
h = ax2.axvspan(times[c.start], times[c.stop - 1],
color='r', alpha=0.3)
else:
ax2.axvspan(times[c.start], times[c.stop - 1], color=(0.3, 0.3, 0.3),
alpha=0.3)
hf = plt.plot(times, T_obs, 'g')
ax2.legend((h, ), ('cluster p-value < 0.05', ))
ax2.set_xlabel("time (ms)")
ax2.set_ylabel("f-values")
Total running time of the script: ( 0 minutes 5.316 seconds)
Estimated memory usage: 9 MB