mne.minimum_norm.apply_inverse_cov#
- mne.minimum_norm.apply_inverse_cov(cov, info, inverse_operator, nave=1, lambda2=0.1111111111111111, method='dSPM', pick_ori=None, prepared=False, label=None, method_params=None, use_cps=True, verbose=None)[source]#
Apply inverse operator to covariance data.
- Parameters:
- covinstance of
Covariance Covariance data, computed on the time segment for which to compute source power.
- info
mne.Info The
mne.Infoobject with information about the sensors and methods of measurement. Used specify the channels to include.- inverse_operatorinstance of
InverseOperator Inverse operator.
- nave
int Number of averages used to regularize the solution.
- lambda2
float The regularization parameter.
- method“MNE” | “dSPM” | “sLORETA” | “eLORETA”
Use minimum norm, dSPM (default), sLORETA, or eLORETA.
- pick_ori
None| “normal” Options:
NonePooling is performed by taking the norm of loose/free orientations. In case of a fixed source space no norm is computed leading to signed source activity.
"normal"Only the normal to the cortical surface is kept. This is only implemented when working with loose orientations.
- preparedbool
If True, do not call
prepare_inverse_operator().- label
Label|None Restricts the source estimates to a given label. If None, source estimates will be computed for the entire source space.
- method_params
dict|None Additional options for eLORETA. See Notes for details.
- use_cpsbool
Whether to use cortical patch statistics to define normal orientations for surfaces (default True).
- verbosebool |
str|int|None Control verbosity of the logging output. If
None, use the default verbosity level. See the logging documentation andmne.verbose()for details. Should only be passed as a keyword argument.
- covinstance of
- Returns:
- stc
SourceEstimate|VectorSourceEstimate|VolSourceEstimate The source estimates.
- stc
See also
apply_inverseApply inverse operator to evoked object.
apply_inverse_rawApply inverse operator to raw object.
apply_inverse_epochsApply inverse operator to epochs object.
apply_inverse_tfr_epochsApply inverse operator to epochs tfr object.
Notes
New in v0.20.
This code is based on the original research code from [1] and has been useful to correct for individual field spread using source localization in the context of predictive modeling.
References
Examples using mne.minimum_norm.apply_inverse_cov#
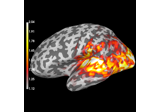
Compute evoked ERS source power using DICS, LCMV beamformer, and dSPM
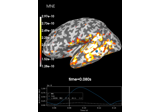
Compute source power estimate by projecting the covariance with MNE