mne.pick_types#
- mne.pick_types(info, meg=False, eeg=False, stim=False, eog=False, ecg=False, emg=False, ref_meg='auto', *, misc=False, resp=False, chpi=False, exci=False, ias=False, syst=False, seeg=False, dipole=False, gof=False, bio=False, ecog=False, fnirs=False, csd=False, dbs=False, temperature=False, gsr=False, eyetrack=False, include=(), exclude='bads', selection=None)[source]#
Pick channels by type and names.
- Parameters:
- info
mne.Info The
mne.Infoobject with information about the sensors and methods of measurement.- megbool |
str If True include MEG channels. If string it can be ‘mag’, ‘grad’, ‘planar1’ or ‘planar2’ to select only magnetometers, all gradiometers, or a specific type of gradiometer.
- eegbool
If True include EEG channels.
- stimbool
If True include stimulus channels.
- eogbool
If True include EOG channels.
- ecgbool
If True include ECG channels.
- emgbool
If True include EMG channels.
- ref_megbool |
str If True include CTF / 4D reference channels. If ‘auto’, reference channels are included if compensations are present and
megis not False. Can also be the string options for themegparameter.- miscbool
If True include miscellaneous analog channels.
- respbool
If
Trueinclude respiratory channels.- chpibool
If True include continuous HPI coil channels.
- excibool
Flux excitation channel used to be a stimulus channel.
- iasbool
Internal Active Shielding data (maybe on Triux only).
- systbool
System status channel information (on Triux systems only).
- seegbool
Stereotactic EEG channels.
- dipolebool
Dipole time course channels.
- gofbool
Dipole goodness of fit channels.
- biobool
Bio channels.
- ecogbool
Electrocorticography channels.
- fnirsbool |
str Functional near-infrared spectroscopy channels. If True include all fNIRS channels. If False (default) include none. If string it can be ‘hbo’ (to include channels measuring oxyhemoglobin) or ‘hbr’ (to include channels measuring deoxyhemoglobin).
- csdbool
EEG-CSD channels.
- dbsbool
Deep brain stimulation channels.
- temperaturebool
Temperature channels.
- gsrbool
Galvanic skin response channels.
- eyetrackbool |
str Eyetracking channels. If True include all eyetracking channels. If False (default) include none. If string it can be ‘eyegaze’ (to include eye position channels) or ‘pupil’ (to include pupil-size channels).
- include
listofstr List of additional channels to include. If empty do not include any.
- exclude
listofstr|str List of channels to exclude. If ‘bads’ (default), exclude channels in
info['bads'].- selection
listofstr Restrict sensor channels (MEG, EEG, etc.) to this list of channel names.
- info
- Returns:
Examples using mne.pick_types#
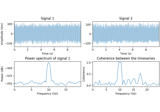
Motor imagery decoding from EEG data using the Common Spatial Pattern (CSP)
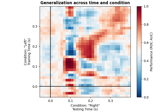

Decoding sensor space data with generalization across time and conditions
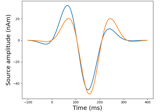
Analysis of evoked response using ICA and PCA reduction techniques
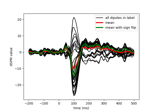

Compute MNE-dSPM inverse solution on single epochs
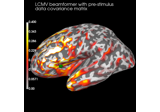
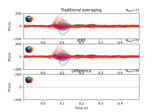
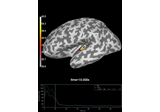
Compute evoked ERS source power using DICS, LCMV beamformer, and dSPM
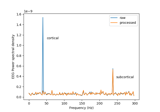
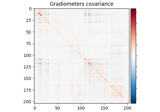
Cortical Signal Suppression (CSS) for removal of cortical signals
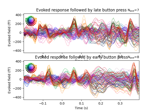
Define target events based on time lag, plot evoked response
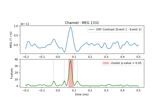

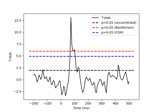
Permutation F-test on sensor data with 1D cluster level
Compute Power Spectral Density of inverse solution from single epochs
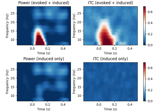

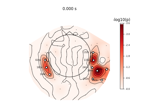
Compute power and phase lock in label of the source space
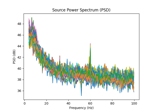
Compute source power spectral density (PSD) in a label
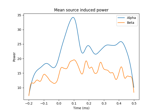
Compute induced power in the source space with dSPM
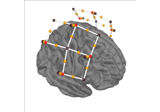
Preprocessing functional near-infrared spectroscopy (fNIRS) data
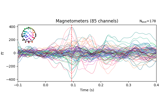
Preprocessing optically pumped magnetometer (OPM) MEG data
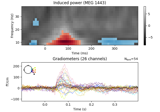
Non-parametric 1 sample cluster statistic on single trial power
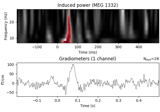
Non-parametric between conditions cluster statistic on single trial power
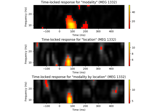

Mass-univariate twoway repeated measures ANOVA on single trial power

Spatiotemporal permutation F-test on full sensor data
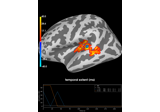
Permutation t-test on source data with spatio-temporal clustering
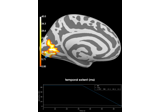
Repeated measures ANOVA on source data with spatio-temporal clustering
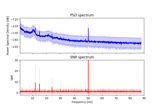

Frequency-tagging: Basic analysis of an SSVEP/vSSR dataset