mne.make_forward_solution#
- mne.make_forward_solution(info, trans, src, bem, meg=True, eeg=True, *, mindist=0.0, ignore_ref=False, n_jobs=None, on_inside='raise', verbose=None)[source]#
Calculate a forward solution for a subject.
- Parameters:
- info
mne.Info| path-like The
mne.Infoobject with information about the sensors and methods of measurement. Ifpath-like, it should be astrorpathlib.Pathto a file with measurement information (e.g.mne.io.Raw).- transpath-like |
dict| instance ofTransform|"fsaverage"|None If str, the path to the head<->MRI transform
*-trans.fiffile produced during coregistration. Can also be'fsaverage'to use the built-in fsaverage transformation. If trans is None, an identity matrix is assumed.Changed in version 0.19: Support for
'fsaverage'argument.- srcpath-like | instance of
SourceSpaces Either a path to a source space file or a loaded or generated
SourceSpaces.- bempath-like |
ConductorModel Filename of the BEM (e.g.,
"sample-5120-5120-5120-bem-sol.fif") to use, or a loadedConductorModel. Seemake_bem_model()andmake_bem_solution()to create amne.bem.ConductorModel.- megbool
If True (default), include MEG computations.
- eegbool
If True (default), include EEG computations.
- mindist
float Minimum distance of sources from inner skull surface (in mm).
- ignore_refbool
If True, do not include reference channels in compensation. This option should be True for KIT files, since forward computation with reference channels is not currently supported.
- n_jobs
int|None The number of jobs to run in parallel. If
-1, it is set to the number of CPU cores. Requires thejoblibpackage.None(default) is a marker for ‘unset’ that will be interpreted asn_jobs=1(sequential execution) unless the call is performed under ajoblib.parallel_configcontext manager that sets another value forn_jobs.- on_inside‘raise’ | ‘warn’ | ‘ignore’
What to do if MEG sensors are inside the outer skin surface. If ‘raise’ (default), an error is raised. If ‘warn’ or ‘ignore’, the forward solution is computed anyway and a warning is or isn’t emitted, respectively.
New in v1.10.
- verbosebool |
str|int|None Control verbosity of the logging output. If
None, use the default verbosity level. See the logging documentation andmne.verbose()for details. Should only be passed as a keyword argument.
- info
- Returns:
- fwdinstance of
Forward The forward solution.
- fwdinstance of
See also
Notes
The
--gradoption from MNE-C (to compute gradients) is not implemented here.To create a fixed-orientation forward solution, use this function followed by
mne.convert_forward_solution().Note
If the BEM solution was computed with OpenMEEG in
mne.make_bem_solution(), then OpenMEEG will automatically be used to compute the forward solution.Changed in version 1.2: Added support for OpenMEEG-based forward solution calculations.
Examples using mne.make_forward_solution#
Compute MNE inverse solution on evoked data with a mixed source space
Compute source power spectral density (PSD) of VectorView and OPM data
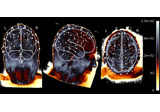
EEG source localization given electrode locations on an MRI
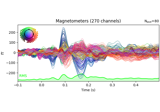
Working with CTF data: the Brainstorm auditory dataset
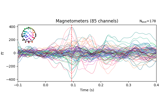
Preprocessing optically pumped magnetometer (OPM) MEG data