mne.combine_evoked#
- mne.combine_evoked(all_evoked, weights)[source]#
Merge evoked data by weighted addition or subtraction.
Each
Evokedinall_evokedshould have the same channels and the same time instants. Subtraction can be performed by passingweights=[1, -1].Warning
Other than cases like simple subtraction mentioned above (where all weights are
-1or1), if you provide numeric weights instead of using'equal'or'nave', the resultingEvokedobject’s.naveattribute (which is used to scale noise covariance when applying the inverse operator) may not be suitable for inverse imaging.- Parameters:
- all_evoked
listofEvoked The evoked datasets.
- weights
listoffloat|'equal'|'nave' The weights to apply to the data of each evoked instance, or a string describing the weighting strategy to apply:
'nave'computes sum-to-one weights proportional to each object’snaveattribute;'equal'weights eachEvokedby1 / len(all_evoked).
- all_evoked
- Returns:
- evoked
Evoked The new evoked data.
- evoked
Notes
New in v0.9.0.
Examples using mne.combine_evoked#
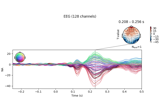
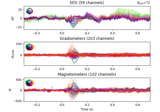
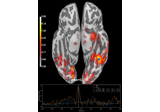
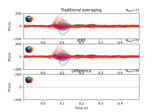
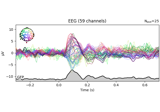
Single trial linear regression analysis with the LIMO dataset
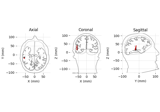
Source localization with equivalent current dipole (ECD) fit
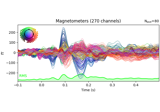
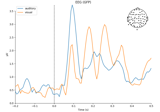
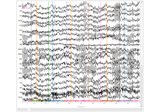
Working with CTF data: the Brainstorm auditory dataset
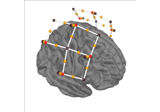
Preprocessing functional near-infrared spectroscopy (fNIRS) data
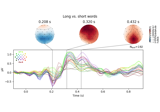
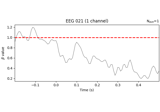
Visualising statistical significance thresholds on EEG data
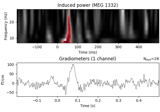
Non-parametric between conditions cluster statistic on single trial power