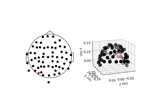
mne.channels.make_standard_montage#
- mne.channels.make_standard_montage(kind, head_size='auto')[source]#
Read a generic (built-in) standard montage that ships with MNE-Python.
- Parameters:
- kind
str The name of the montage to use.
Note
You can retrieve the names of all built-in montages via
mne.channels.get_builtin_montages().- head_size
float|None|str The head size (radius, in meters) to use for spherical montages. Can be None to not scale the read sizes.
'auto'(default) will use 95mm for all montages except the'colin27*','mgh*', and'artinis*', which are already in fsaverage’s MRI coordinates (same as MNI).
- kind
- Returns:
- montageinstance of
DigMontage The montage.
- montageinstance of
Notes
Individualized (digitized) electrode positions should be read in using
read_dig_captrak(),read_dig_curry(),read_dig_egi(),read_dig_fif(),read_dig_polhemus_isotrak(),read_dig_hpts(), or manually made withmake_dig_montage().New in v0.19.0.
Examples using mne.channels.make_standard_montage#
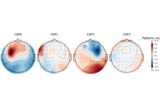
Motor imagery decoding from EEG data using the Common Spatial Pattern (CSP)
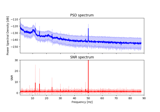

Frequency-tagging: Basic analysis of an SSVEP/vSSR dataset