mne.Forward#
- class mne.Forward[source]#
Forward class to represent info from forward solution.
Like
mne.Info, this data structure behaves like a dictionary. It contains all metadata necessary for a forward solution.Warning
This class should not be modified or created by users. Forward objects should be obtained using
mne.make_forward_solution()ormne.read_forward_solution().- Attributes:
Methods
copy()Copy the Forward instance.
pick_channels(ch_names[, ordered])Pick channels from this forward operator.
save(fname, *[, overwrite, verbose])Save the forward solution.
Notes
Forward data is accessible via string keys using standard
dictaccess (e.g.,fwd['nsource'] == 4096):- source_oriint
The source orientation, either
FIFF.FIFFV_MNE_FIXED_ORIorFIFF.FIFFV_MNE_FREE_ORI.- coord_frameint
The coordinate frame of the forward solution, usually
FIFF.FIFFV_COORD_HEAD.- nsourceint
The number of source locations.
- nchanint
The number of channels.
- soldict
The forward solution, with entries:
'data'ndarray, shape (n_channels, nsource * n_ori)The forward solution data. The shape will be
(n_channels, nsource)for a fixed-orientation forward and(n_channels, nsource * 3)for a free-orientation forward.'row_names'list of strThe channel names.
- mri_head_tinstance of Transform
The mri ↔ head transformation that was used.
- infoinstance of
Info The measurement information (with contents reduced compared to that of the original data).
- srcinstance of
SourceSpaces The source space used during forward computation. This can differ from the original source space as:
Source points are removed due to proximity to (or existing outside) the inner skull surface.
The source space will be converted to the
coord_frameof the forward solution, which typically means it gets converted from MRI to head coordinates.
- source_rrndarray, shape (n_sources, 3)
The source locations.
- source_nnndarray, shape (n_sources, 3)
The source normals. Will be all +Z (
(0, 0, 1.)) for volume source spaces. For surface source spaces, these are normal to the cortical surface.- surf_oriint
Whether
solis surface-oriented with the surface normal in the Z component (FIFF.FIFFV_MNE_FIXED_ORI) or +Z in the givencoord_framein the Z component (FIFF.FIFFV_MNE_FREE_ORI).
Forward objects also have some attributes that are accessible via
.access, likefwd.ch_names.- pick_channels(ch_names, ordered=False)[source]#
Pick channels from this forward operator.
- Parameters:
- Returns:
- fwdinstance of Forward.
The modified forward model.
Notes
Operates in-place.
New in v0.20.0.
- save(fname, *, overwrite=False, verbose=None)[source]#
Save the forward solution.
- Parameters:
- fnamepath-like
File name to save the forward solution to. It should end with
-fwd.fifor-fwd.fif.gzto save to FIF, or-fwd.h5to save to HDF5.- overwritebool
If True (default False), overwrite the destination file if it exists.
- verbosebool |
str|int|None Control verbosity of the logging output. If
None, use the default verbosity level. See the logging documentation andmne.verbose()for details. Should only be passed as a keyword argument.
Examples using mne.Forward#
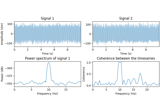
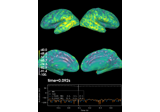
Compute source level time-frequency timecourses using a DICS beamformer
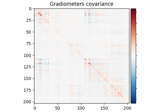
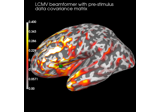
Compute evoked ERS source power using DICS, LCMV beamformer, and dSPM
Compute a sparse inverse solution using the Gamma-MAP empirical Bayesian method

Compute sparse inverse solution with mixed norm: MxNE and irMxNE
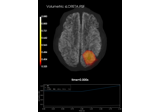
Compute source power estimate by projecting the covariance with MNE

Computing source timecourses with an XFit-like multi-dipole model

Compute iterative reweighted TF-MxNE with multiscale time-frequency dictionary
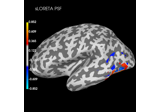
Plot point-spread functions (PSFs) and cross-talk functions (CTFs)
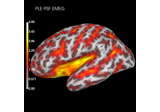
Compute spatial resolution metrics to compare MEG with EEG+MEG
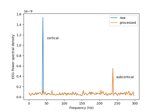
Cortical Signal Suppression (CSS) for removal of cortical signals
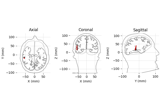
Source localization with equivalent current dipole (ECD) fit
The role of dipole orientations in distributed source localization
EEG source localization given electrode locations on an MRI
Preprocessing optically pumped magnetometer (OPM) MEG data