mne.read_events#
- mne.read_events(filename, include=None, exclude=None, mask=None, mask_type='and', return_event_id=False, verbose=None)[source]#
Read events from fif or text file.
See Parsing events from raw data and Working with events for more information about events.
- Parameters:
- filenamepath-like
Name of the input file. If the extension is
.fif, events are read assuming the file is in FIF format, otherwise (e.g.,.eve,.lst,.txt) events are read as coming from text. Note that new format event files do not contain the"time"column (used to be the second column).- include
int|list|None A event id to include or a list of them. If None all events are included.
- exclude
int|list|None A event id to exclude or a list of them. If None no event is excluded. If include is not None the exclude parameter is ignored.
- mask
int|None The value of the digital mask to apply to the stim channel values. If None (default), no masking is performed.
- mask_type
'and'|'not_and' The type of operation between the mask and the trigger. Choose ‘and’ (default) for MNE-C masking behavior.
New in v0.13.
- return_event_idbool
If True,
event_idwill be returned. This is only possible for-annot.fiffiles produced with MNE-Cmne_browse_raw.New in v0.20.
- verbosebool |
str|int|None Control verbosity of the logging output. If
None, use the default verbosity level. See the logging documentation andmne.verbose()for details. Should only be passed as a keyword argument.
- Returns:
See also
Notes
This function will discard the offset line (i.e., first line with zero event number) if it is present in a text file.
For more information on
maskandmask_type, seemne.find_events().
Examples using mne.read_events#

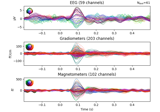
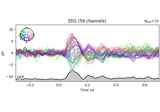
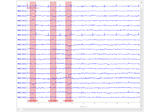
Decoding sensor space data with generalization across time and conditions
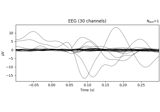
Analysis of evoked response using ICA and PCA reduction techniques
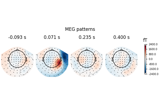

Linear classifier on sensor data with plot patterns and filters

Compute MNE-dSPM inverse solution on single epochs
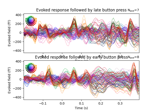
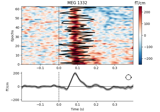
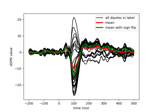
Define target events based on time lag, plot evoked response
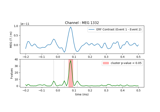

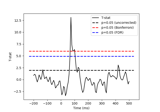
Permutation F-test on sensor data with 1D cluster level
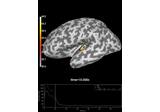
Compute Power Spectral Density of inverse solution from single epochs
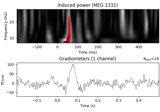
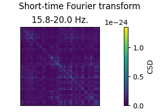
Non-parametric between conditions cluster statistic on single trial power
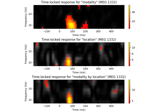

Mass-univariate twoway repeated measures ANOVA on single trial power

Spatiotemporal permutation F-test on full sensor data
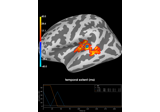
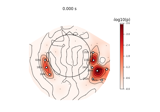
Permutation t-test on source data with spatio-temporal clustering
Repeated measures ANOVA on source data with spatio-temporal clustering