mne.io.read_raw_edf#
- mne.io.read_raw_edf(input_fname, eog=None, misc=None, stim_channel='auto', exclude=(), infer_types=False, include=None, preload=False, units=None, encoding='utf8', exclude_after_unique=False, *, verbose=None) RawEDF[source]#
Reader function for EDF and EDF+ files.
- Parameters:
- input_fnamepath-like
Path to the EDF or EDF+ file or EDF/EDF+ file itself. If a file-like object is provided, preload must be used.
Changed in version 1.10: Added support for file-like objects
- eog
listortuple Names of channels or list of indices that should be designated EOG channels. Values should correspond to the electrodes in the file. Default is None.
- misc
listortuple Names of channels or list of indices that should be designated MISC channels. Values should correspond to the electrodes in the file. Default is None.
- stim_channel
'auto'|str|listofstr|int|listofint Defaults to
'auto', which means that channels named'status'or'trigger'(case insensitive) are set to STIM. If str (or list of str), all channels matching the name(s) are set to STIM. If int (or list of ints), channels corresponding to the indices are set to STIM.- exclude
listofstr|str Channel names to exclude. This can help when reading data with different sampling rates to avoid unnecessary resampling. A str is interpreted as a regular expression.
- infer_typesbool
If True, try to infer channel types from channel labels. If a channel label starts with a known type (such as ‘EEG’) followed by a space and a name (such as ‘Fp1’), the channel type will be set accordingly, and the channel will be renamed to the original label without the prefix. For unknown prefixes, the type will be ‘EEG’ and the name will not be modified. If False, do not infer types and assume all channels are of type ‘EEG’.
New in v0.24.1.
- include
listofstr|str Channel names to be included. A str is interpreted as a regular expression. ‘exclude’ must be empty if include is assigned.
New in v1.1.
- preloadbool or
str(defaultFalse) Preload data into memory for data manipulation and faster indexing. If True, the data will be preloaded into memory (fast, requires large amount of memory). If preload is a string, preload is the file name of a memory-mapped file which is used to store the data on the hard drive (slower, requires less memory).
- units
dict|str The units of the channels as stored in the file. This argument is useful only if the units are missing from the original file. If a dict, it must map a channel name to its unit, and if str it is assumed that all channels have the same units.
- encoding
str Encoding of annotations channel(s). Default is “utf8” (the only correct encoding according to the EDF+ standard).
- exclude_after_uniquebool
If True, exclude channels are searched for after they have been made unique. This is useful to choose channels that have been made unique by adding a suffix. If False, the original names are checked.
Changed in version 1.7.
- verbosebool |
str|int|None Control verbosity of the logging output. If
None, use the default verbosity level. See the logging documentation andmne.verbose()for details. Should only be passed as a keyword argument.
- Returns:
- rawinstance of
RawEDF The raw instance. See
mne.io.Rawfor documentation of attributes and methods.
- rawinstance of
See also
mne.io.read_raw_bdfReader function for BDF files.
mne.io.read_raw_gdfReader function for GDF files.
mne.export.export_rawExport function for EDF files.
mne.io.RawDocumentation of attributes and methods of RawEDF.
Notes
mne.io.Rawonly stores signals with matching sampling frequencies. Therefore, if mixed sampling frequency signals are requested, all signals are upsampled to the highest loaded sampling frequency. In this case, using preload=True is recommended, as otherwise, edge artifacts appear when slices of the signal are requested.It is worth noting that in some special cases, it may be necessary to shift event values in order to retrieve correct event triggers. This depends on the triggering device used to perform the synchronization. For instance, in some files events need to be shifted by 8 bits:
>>> events[:, 2] >>= 8
TAL channels called ‘EDF Annotations’ are parsed and extracted annotations are stored in raw.annotations. Use
mne.events_from_annotations()to obtain events from these annotations.If channels named ‘status’ or ‘trigger’ are present, they are considered as STIM channels by default. Use
mne.find_events()to parse events encoded in such analog stim channels.The EDF specification allows optional storage of channel types in the prefix of the signal label for each channel. For example,
EEG Fzimplies thatFzis an EEG channel andMISC Ewould implyEis a MISC channel. However, there is no standard way of specifying all channel types. MNE-Python will try to infer the channel type, when such a string exists, defaulting to EEG, when there is no prefix or the prefix is not recognized.The following prefix strings are mapped to MNE internal types:
‘EEG’: ‘eeg’
‘SEEG’: ‘seeg’
‘ECOG’: ‘ecog’
‘DBS’: ‘dbs’
‘EOG’: ‘eog’
‘ECG’: ‘ecg’
‘EMG’: ‘emg’
‘BIO’: ‘bio’
‘RESP’: ‘resp’
‘MISC’: ‘misc’
‘SAO2’: ‘bio’
The EDF specification allows storage of subseconds in measurement date. However, this reader currently sets subseconds to 0 by default.
Examples using mne.io.read_raw_edf#
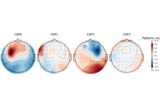
Motor imagery decoding from EEG data using the Common Spatial Pattern (CSP)

Decoding in time-frequency space using Common Spatial Patterns (CSP)
Sleep stage classification from polysomnography (PSG) data