Note
Go to the end to download the full example code.
2 samples permutation test on source data with spatio-temporal clustering#
Tests if the source space data are significantly different between 2 groups of subjects (simulated here using one subject’s data). The multiple comparisons problem is addressed with a cluster-level permutation test across space and time.
# Authors: Alexandre Gramfort <alexandre.gramfort@inria.fr>
# Eric Larson <larson.eric.d@gmail.com>
# License: BSD-3-Clause
# Copyright the MNE-Python contributors.
import numpy as np
from scipy import stats as stats
import mne
from mne import spatial_src_adjacency
from mne.datasets import sample
from mne.stats import spatio_temporal_cluster_test, summarize_clusters_stc
print(__doc__)
Set parameters#
data_path = sample.data_path()
meg_path = data_path / "MEG" / "sample"
stc_fname = meg_path / "sample_audvis-meg-lh.stc"
subjects_dir = data_path / "subjects"
src_fname = subjects_dir / "fsaverage" / "bem" / "fsaverage-ico-5-src.fif"
# Load stc to in common cortical space (fsaverage)
stc = mne.read_source_estimate(stc_fname)
stc.resample(50, npad="auto")
# Read the source space we are morphing to
src = mne.read_source_spaces(src_fname)
fsave_vertices = [s["vertno"] for s in src]
morph = mne.compute_source_morph(
stc,
"sample",
"fsaverage",
spacing=fsave_vertices,
smooth=20,
subjects_dir=subjects_dir,
)
stc = morph.apply(stc)
n_vertices_fsave, n_times = stc.data.shape
tstep = stc.tstep * 1000 # convert to milliseconds
n_subjects1, n_subjects2 = 6, 7
print(f"Simulating data for {n_subjects1} and {n_subjects2} subjects.")
# Let's make sure our results replicate, so set the seed.
np.random.seed(0)
X1 = np.random.randn(n_vertices_fsave, n_times, n_subjects1) * 10
X2 = np.random.randn(n_vertices_fsave, n_times, n_subjects2) * 10
X1[:, :, :] += stc.data[:, :, np.newaxis]
# make the activity bigger for the second set of subjects
X2[:, :, :] += 3 * stc.data[:, :, np.newaxis]
# We want to compare the overall activity levels for each subject
X1 = np.abs(X1) # only magnitude
X2 = np.abs(X2) # only magnitude
Reading a source space...
[done]
Reading a source space...
[done]
2 source spaces read
surface source space present ...
Computing morph matrix...
Left-hemisphere map read.
Right-hemisphere map read.
20 smooth iterations done.
20 smooth iterations done.
[done]
[done]
Simulating data for 6 and 7 subjects.
Compute statistic#
To use an algorithm optimized for spatio-temporal clustering, we just pass the spatial adjacency matrix (instead of spatio-temporal)
print("Computing adjacency.")
adjacency = spatial_src_adjacency(src)
# Note that X needs to be a list of multi-dimensional array of shape
# samples (subjects_k) × time × space, so we permute dimensions
X1 = np.transpose(X1, [2, 1, 0])
X2 = np.transpose(X2, [2, 1, 0])
X = [X1, X2]
# Now let's actually do the clustering. This can take a long time...
# Here we set the threshold quite high to reduce computation,
# and use a very low number of permutations for the same reason.
n_permutations = 50
p_threshold = 0.001
f_threshold = stats.distributions.f.ppf(
1.0 - p_threshold / 2.0, n_subjects1 - 1, n_subjects2 - 1
)
print("Clustering.")
F_obs, clusters, cluster_p_values, H0 = clu = spatio_temporal_cluster_test(
X,
adjacency=adjacency,
n_jobs=None,
n_permutations=n_permutations,
threshold=f_threshold,
buffer_size=None,
)
# Now select the clusters that are sig. at p < 0.05 (note that this value
# is multiple-comparisons corrected).
good_cluster_inds = np.where(cluster_p_values < 0.05)[0]
Computing adjacency.
-- number of adjacent vertices : 20484
Clustering.
stat_fun(H1): min=1.8724146338349605e-13 max=303.63217183709247
Running initial clustering …
Found 361 clusters
0%| | Permuting : 0/49 [00:00<?, ?it/s]
2%|▏ | Permuting : 1/49 [00:00<00:01, 29.62it/s]
6%|▌ | Permuting : 3/49 [00:00<00:01, 44.85it/s]
10%|█ | Permuting : 5/49 [00:00<00:00, 49.90it/s]
12%|█▏ | Permuting : 6/49 [00:00<00:00, 44.41it/s]
16%|█▋ | Permuting : 8/49 [00:00<00:00, 46.84it/s]
20%|██ | Permuting : 10/49 [00:00<00:00, 49.16it/s]
22%|██▏ | Permuting : 11/49 [00:00<00:00, 45.98it/s]
24%|██▍ | Permuting : 12/49 [00:00<00:00, 43.13it/s]
29%|██▊ | Permuting : 14/49 [00:00<00:00, 44.28it/s]
33%|███▎ | Permuting : 16/49 [00:00<00:00, 46.09it/s]
35%|███▍ | Permuting : 17/49 [00:00<00:00, 43.58it/s]
39%|███▉ | Permuting : 19/49 [00:00<00:00, 45.22it/s]
41%|████ | Permuting : 20/49 [00:00<00:00, 43.69it/s]
45%|████▍ | Permuting : 22/49 [00:00<00:00, 45.17it/s]
47%|████▋ | Permuting : 23/49 [00:00<00:00, 43.78it/s]
49%|████▉ | Permuting : 24/49 [00:00<00:00, 42.56it/s]
53%|█████▎ | Permuting : 26/49 [00:00<00:00, 43.97it/s]
55%|█████▌ | Permuting : 27/49 [00:00<00:00, 42.81it/s]
59%|█████▉ | Permuting : 29/49 [00:00<00:00, 44.11it/s]
61%|██████ | Permuting : 30/49 [00:00<00:00, 43.02it/s]
65%|██████▌ | Permuting : 32/49 [00:00<00:00, 44.23it/s]
67%|██████▋ | Permuting : 33/49 [00:00<00:00, 43.18it/s]
69%|██████▉ | Permuting : 34/49 [00:00<00:00, 42.22it/s]
73%|███████▎ | Permuting : 36/49 [00:00<00:00, 43.39it/s]
78%|███████▊ | Permuting : 38/49 [00:00<00:00, 44.48it/s]
80%|███████▉ | Permuting : 39/49 [00:00<00:00, 43.43it/s]
84%|████████▎ | Permuting : 41/49 [00:00<00:00, 44.49it/s]
86%|████████▌ | Permuting : 42/49 [00:00<00:00, 43.53it/s]
90%|████████▉ | Permuting : 44/49 [00:00<00:00, 44.54it/s]
92%|█████████▏| Permuting : 45/49 [00:01<00:00, 43.58it/s]
96%|█████████▌| Permuting : 47/49 [00:01<00:00, 44.56it/s]
100%|██████████| Permuting : 49/49 [00:01<00:00, 45.52it/s]
100%|██████████| Permuting : 49/49 [00:01<00:00, 44.77it/s]
Visualize the clusters#
print("Visualizing clusters.")
# Now let's build a convenient representation of each cluster, where each
# cluster becomes a "time point" in the SourceEstimate
fsave_vertices = [np.arange(10242), np.arange(10242)]
stc_all_cluster_vis = summarize_clusters_stc(
clu, tstep=tstep, vertices=fsave_vertices, subject="fsaverage"
)
# Let's actually plot the first "time point" in the SourceEstimate, which
# shows all the clusters, weighted by duration
# blue blobs are for condition A != condition B
brain = stc_all_cluster_vis.plot(
"fsaverage",
hemi="both",
views="lateral",
subjects_dir=subjects_dir,
time_label="temporal extent (ms)",
clim=dict(kind="value", lims=[0, 1, 40]),
)
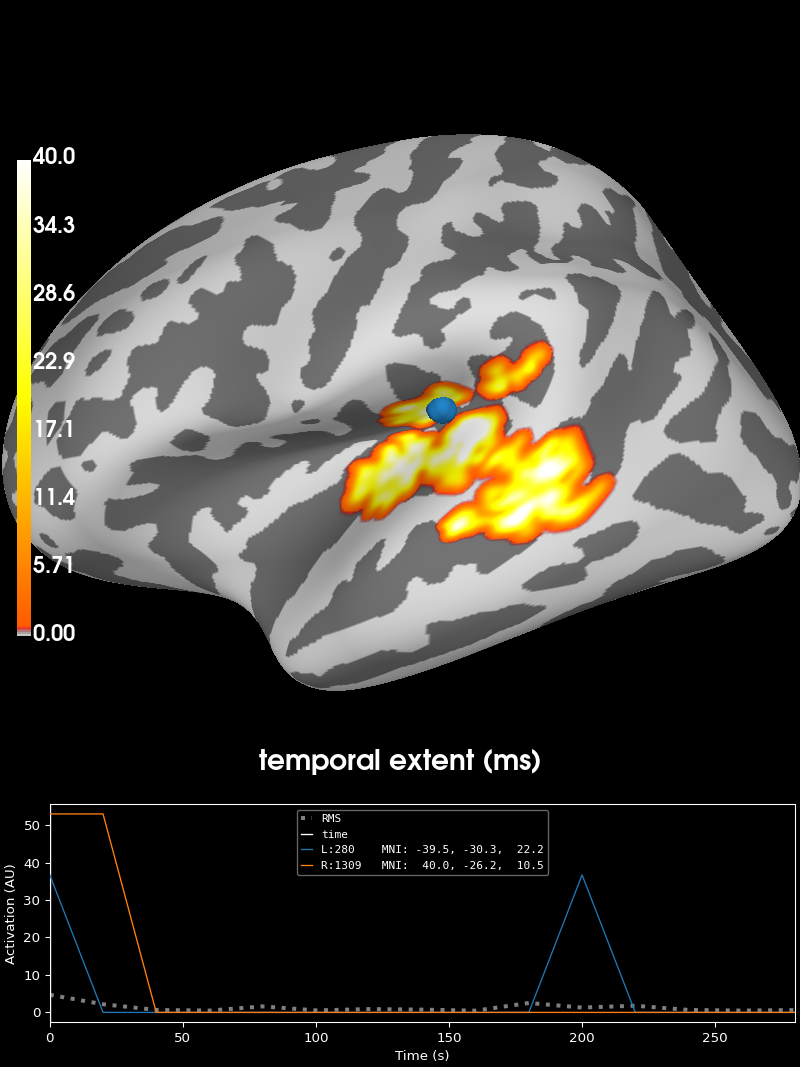
Visualizing clusters.
Total running time of the script: (0 minutes 10.759 seconds)