Note
Go to the end to download the full example code.
Reading BCI2000 files#
In this example, we use MNE-Python to read a BCI2000 .dat file.
BCI2000 is a general-purpose brain-computer interface (BCI) system widely
used in EEG research. The file is downloaded from the MNE testing data
repository using pooch.
# Authors: The MNE-Python contributors.
# License: BSD-3-Clause
# Copyright the MNE-Python contributors.
import pooch
import mne
First, we download the sample BCI2000 .dat file using pooch.
data_dir = mne.datasets.default_path() / "bci2k_data"
data_dir.mkdir(exist_ok=True)
fname = pooch.retrieve(
url="https://raw.githubusercontent.com/mne-tools/mne-testing-data/master/BCI2k/bci2k_test.dat",
known_hash="sha256:8efc7b5f700660a044086cb1449806ca408c2e6d32d9338c32e1bf31ce3ca9cb",
path=data_dir,
)
Using default location ~/mne_data...
Now we can read the file using mne.io.read_raw_bci2k().
Note that preload=True is required for BCI2000 files.
raw = mne.io.read_raw_bci2k(fname, preload=True)
print(raw.info)
<Info | 7 non-empty values
bads: []
ch_names: EEG1, EEG2, STI 014
chs: 2 EEG, 1 Stimulus
custom_ref_applied: False
highpass: 0.0 Hz
lowpass: 128.0 Hz
meas_date: unspecified
nchan: 3
projs: []
sfreq: 256.0 Hz
>
We can inspect the object representation, channel names, types, sampling frequency, and recording duration.
print(raw)
print(f"Channel names : {raw.ch_names}")
print(f"Channel types : {raw.get_channel_types()}")
print(f"Sampling freq : {raw.info['sfreq']} Hz")
print(f"Duration : {raw.times[-1]:.2f} s")
print(f"n_channels : {raw.info['nchan']}")
print(f"Data shape : {raw.get_data().shape} (n_channels, n_samples)")
<RawBCI2k | e5d7337a8476d2c2629d5156efe5788c-bci2k_test.dat, 3 x 9248 (36.1 s), ~223 KiB, data loaded>
Channel names : ['EEG1', 'EEG2', 'STI 014']
Channel types : ['eeg', 'eeg', 'stim']
Sampling freq : 256.0 Hz
Duration : 36.12 s
n_channels : 3
Data shape : (3, 9248) (n_channels, n_samples)
If the BCI2000 file contains a StimulusCode state, it is automatically
mapped to a STI 014 stim channel. We can extract events from it using
mne.find_events().
if "STI 014" in raw.ch_names:
events = mne.find_events(raw, shortest_event=1)
print(f"Found {len(events)} events")
print(mne.count_events(events))
else:
print("No stim channel found in this file.")
Finding events on: STI 014
Found 0 events
{}
Finally, we can visualize the raw data.
raw.plot(duration=5, n_channels=len(raw.ch_names), scalings="auto")
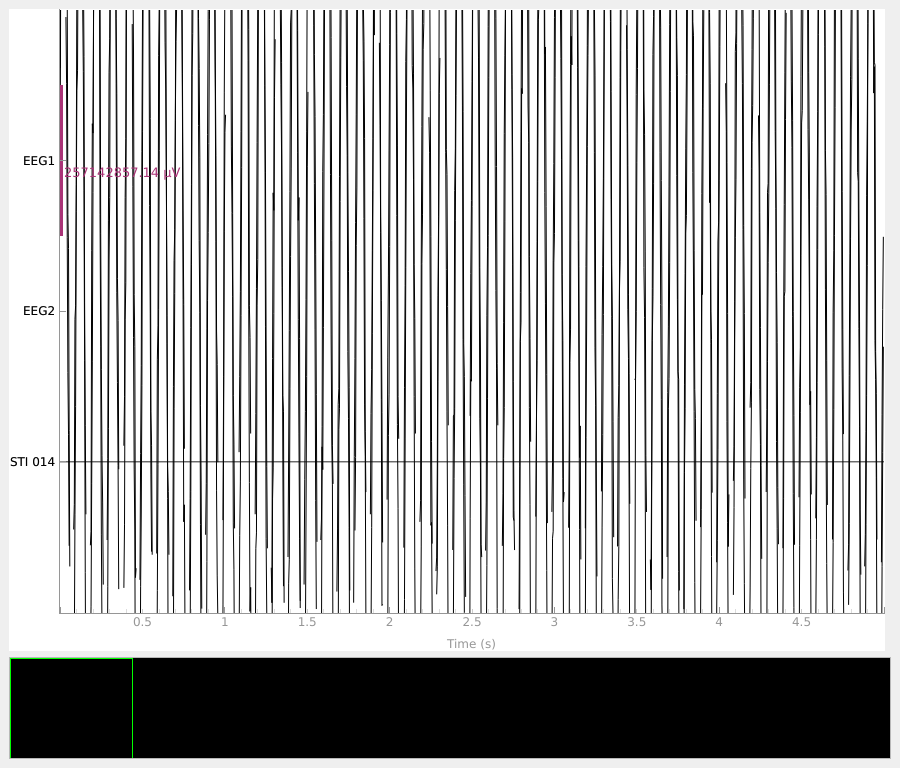
Using qt as 2D backend.
Total running time of the script: (0 minutes 1.582 seconds)